DMD Pattern Illuminator
for Targeted Photostimulation
for Targeted Photostimulation
Polygon
Overview/Features
650+ Labs Worldwide, 130+ Publications Using Polygon
Mightex’s market-leading Polygon DMD pattern illuminator provides precise spatiotemporal control of light with subcellular resolution, making it the perfect illumination tool for life science research.
Cellular-Resolution Optogenetics and Photostimulation
Simultaneous Multi-Region Illumination
Subcellular Resolution
Compatible with Any Microscope

State-of-the-Art Illumination Technology
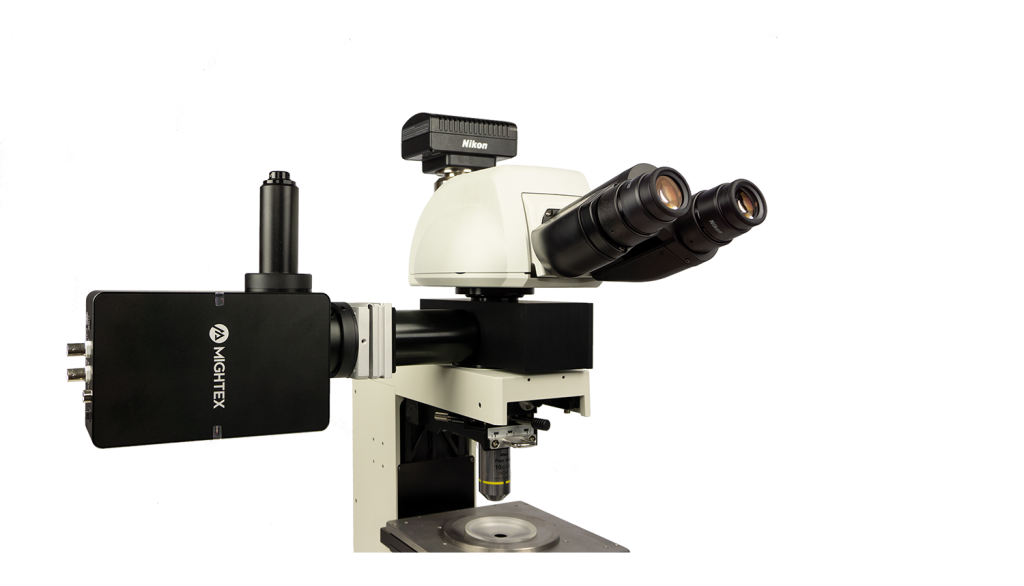
The Polygon uses digital mirror device (DMD) technology to illuminate multiple regions simultaneously. A DMD is composed of hundreds of thousands of micro-mirrors that can be individually turned on to reflect light onto the sample. Thus, you can control each mirror to control the area(s) of illumination and create any number of different sized patterns. The Polygon DMD pattern illuminator can be mounted into the infinity-path of any microscope.
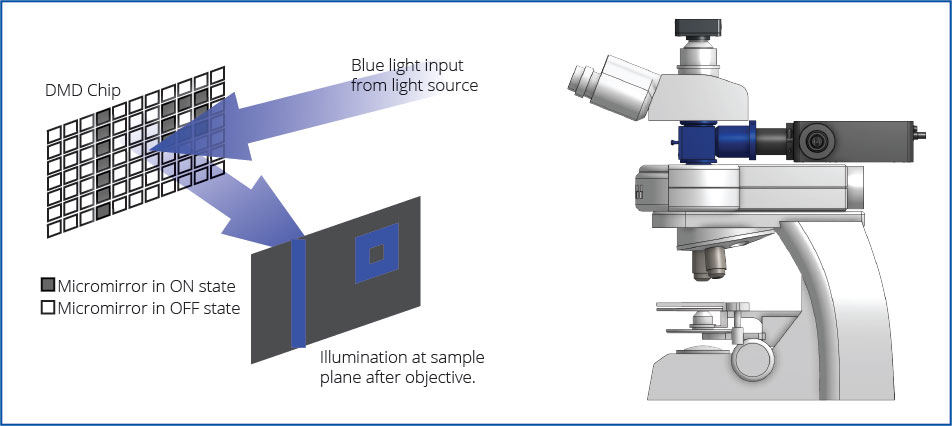
Polygon Key Features
Select illumination of individual cells expressing ChR2 in slice using the Polygon (Courtesy of Matthew Tran & Dr. Blake Richards, McGill University).
Simultaneous Optogenetic Stimulation of Individual Cells or Sub-Cellular Components
The Polygon enables scientists to precisely control where light will hit their sample. With subcellular resolution, the Polygon can illuminate any cellular feature in any shape or size simultaneously on their sample.
Stimulation of Any Optogenetics or Photostimulation Probe
The Polygon provides great flexibility when it comes down to wavelength selection. From UV to VIS/NIR range, the Polygon can project light of different colours suitable for your light-sensitive constructs.
Select gradient illumination of a cell expressing a novel optogenetic construct using the Polygon (Courtesy of Elliot Dine & Dr. Jared Toettcher, Princeton University).
Spatio-Temporal and Intensity Control of Light
Control the initiation, duration, and intensity of light stimulation patterns using the Polygon. Create different waveforms to control the light intensity and duration outputted from the Polygon.
In Vitro and In Vivo Optogenetic Stimulation
The Polygon can be used with a wide-range light sources, including LEDs and lasers. The Polygon is also compatible with Mightex’s OASIS Implant for freely-behaving imaging and optogenetics and OASIS Macro for cortex-wide imaging and optogenetics.
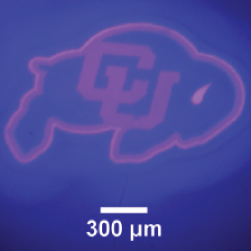

UV Photopatterning (Courtesy of Gayla Berg, University of Colorado).
UV Micropatterning with Sub-Micron Resolution
The Polygon is compatible with UV light sources (350nm+) and provides sub-micron illumination for photopatterning and microfabrication experiments.
Integrates into Any Electrophysiology, Two-Photon or Confocal Microscope Setup
The Polygon is designed to be coupled into the infinity space of any microscope model (Leica, Nikon, Olympus, Zeiss) with Mightex’s microscope-specific adapters. The Polygon comes with BNC connectorized TTL trigger input and output for easy synchronization with different lab equipment.
Illumination patterns using the Polygon synchronized with slice electrophysiological recordings (Courtesy of Matthew Tran and Dr. Blake Richards, McGill University).
Powerful User-Friendly Software for Life Science Experiments
Mightex’s PolyScan software platform is bundled with every Polygon to help you execute sophisticated patterned illumination experiments for your research.
Webinars
650+ Worldwide Polygon Customers
Below is a select list of Universities that are using the Polygon.














130+ Publications Using the Polygon
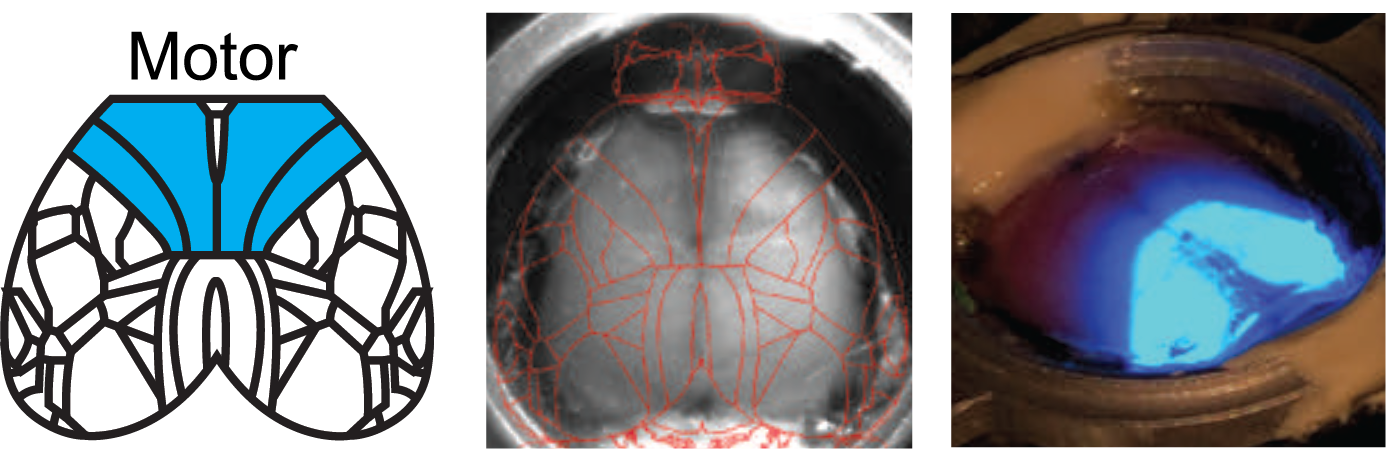
Targeted Optogenetic Stimulation of Motor and Non-Motor Cortices
Kauvar et al. used the Mightex Polygon to optogenetically inhibit the activity of multiple cortical regions—at the same time—in awake mice. They designed a calibration system to align the illumination to specific regions of cortex, and could synchronize illumination to different parts of a task the mouse was performing. This calibration system is demonstrated in these images. The Mightex system enabled them to target nearly the entirety of dorsal cortex.
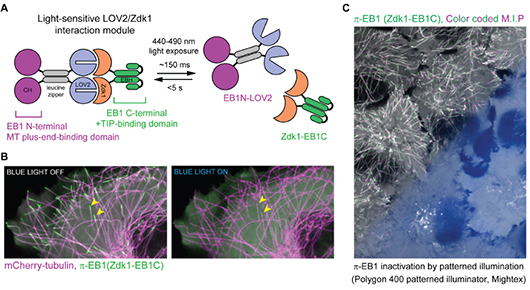
Spatial Control of Microtubules at the Subcellular Scale
van Haren et al. 2018 illustrated the development of a novel optogenetic system to manipulate microtubule organization. Targeted light stimulation from the Polygon was used to precisely inactivate the microtubule-binding protein End-binding protein 1 (EB1), which is responsible for binding microtubules and recruiting plus-end-tracking proteins (+TIPs) to their positive ends.
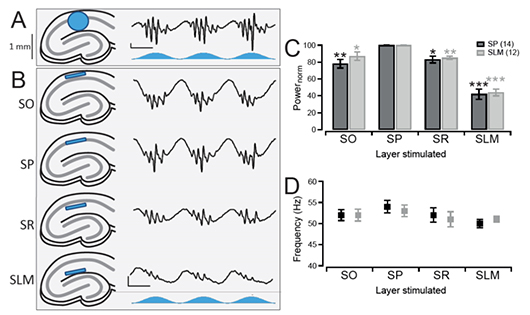
Layer-Specific Optogenetics in the Hippocampus
Butler et al. 2016 explored intrinsic production of gamma oscillations in CA1 region of the hippocampus. Slice electrophysiology was used to measure hippocampal activity, and ChR2 was expressed to manipulate network activity. Mightex’s Polygon was used to illuminate select layers of the hippocampus expressing ChR2 to measure the oscillatory activity produced.
Customer Cases
Customer Cases
With over 650 systems installed in research labs all over the world, Mightex’s market-leading Polygon Pattern Illuminator has enabled scientists to conduct cutting-edge research on a wide range of frontiers such as neuroscience optogenetics, cell biology optogenetics, photoactivation & photoconversion, photopatterning, and microfluidics etc.
Neuroscience Optogenetics
Is it possible to undo cell death in the brain? And can neural functions that are lost as a result of brain damage be regained? Tang et al (2021) explored this topic in their paper Restoration of Visual Function and Cortical Connectivity After Ischemic Injury Through NeuroD1-Mediated Gene Therapy.
For more details, please click here
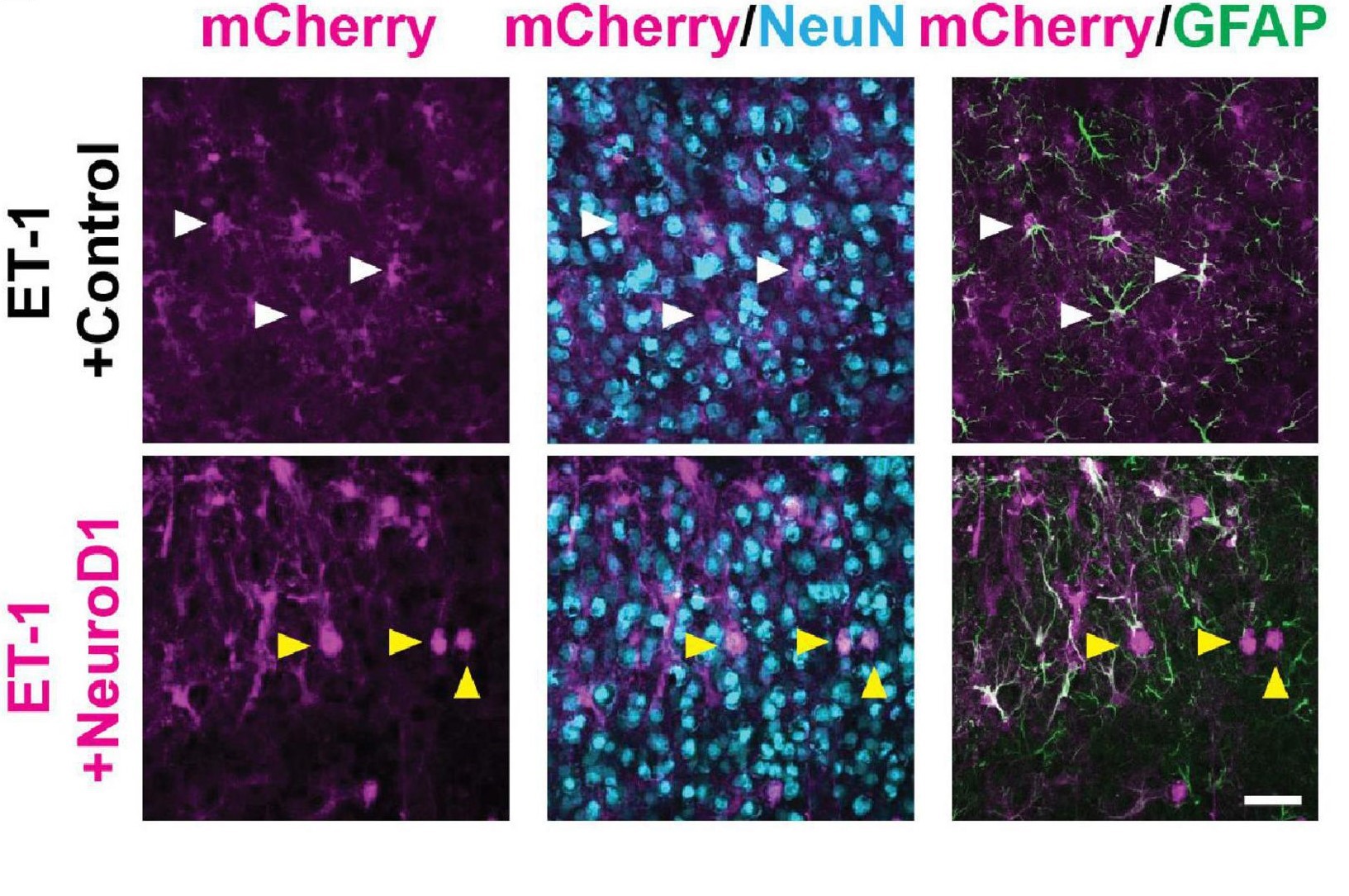
Comparison of cell specific markers in astrocytes. White arrows represent cells co-expressing mCherry and GFAP. Yellow arrows represent cells co-expressing mCherry and Neun. (Courtesy of Dr. Yu Tang & Prof. Alexander Chubykin, Purdue University).
Select illumination of individual cells expressing ChR2 in slice using the Polygon (Courtesy of Matthew Tran & Dr. Blake Richards, McGill University).
Neuroscience Optogenetics
Optogenetics has advanced the field of neuroscience by providing a precise method for manipulating neuronal activity. The ability to perform cellular-resolution optogenetics enables researchers to stimulate select optogenetic-expressing neurons (e.g. ChR2, NpHr, Chrimson etc.) in slice or culture during electrophysiological recordings or calcium imaging to map neural circuitry.
Relevant Publications Using Polygon:
Cell Biology Optogenetics
Optogenetic toolkits have expanded to offer cell biologists unprecedented spatiotemporal molecular and cellular control. From cell migration to gene regulation or developmental processes, cell biologists now have the ability to elicit specific biological responses with an incredible amount of precision in specific cells within organisms or subcellular regions of individual cells using optogenetics.
Select gradient illumination of a cell expressing a novel optogenetic construct using the Polygon (Courtesy of Elliot Dine & Dr. Jared Toettcher, Princeton University).
Relevant Publications Using Polygon:
Johnson HE et al. (2020). Optogenetic Rescue of a Patterning Mutant. Current Biology.
Photoactivation/Photoconversion
Advances in fluorescent protein technology have made photoactivation techniques broadly available in the cell biology lab. By using these techniques, researchers can fluorescently highlight/label specific protein populations in the cell such that their movement and behavior can be tracked over time. Also, photoactivation methods can be applied at different scales, allowing researchers to track proteins within cells, or alternatively to track cells within tissues.
Relevant Publications Using Polygon:
Photopatterning
Many techniques have been developed to manipulate the different properties of the cellular microenviroment. Recently, the advent of light-induced biochemical compounds has allowed scientists to define the biochemical and structural features of cellular matrices by photo micropatterning and photolithography. Scientists can precisely define the spatial distribution of UV light to create unique cellular microenvironments without any use of masks or physical contact with the substrate.

UV Photopatterning (Courtesy of Gayla Berg, University of Colorado).
Relevant Publications Using Polygon:
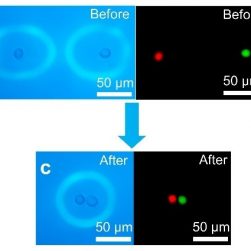

Optoeletronic tweezers to move specific cells (Courtesy of Dr. Shuailong Zhang, University of Toronto).
Microfluidics
Micro-Fluidic devices are made by photo-patterning thin film precursor materials on a substrate to create microscopic channels, chambers, and even flow-controlling mechanisms. Small amounts of fluids containing reactive chemicals or other substances can then flow through the micro-channels and mix in the micro-chambers of the device and react with other substances that may be deposited in the micro-channels.
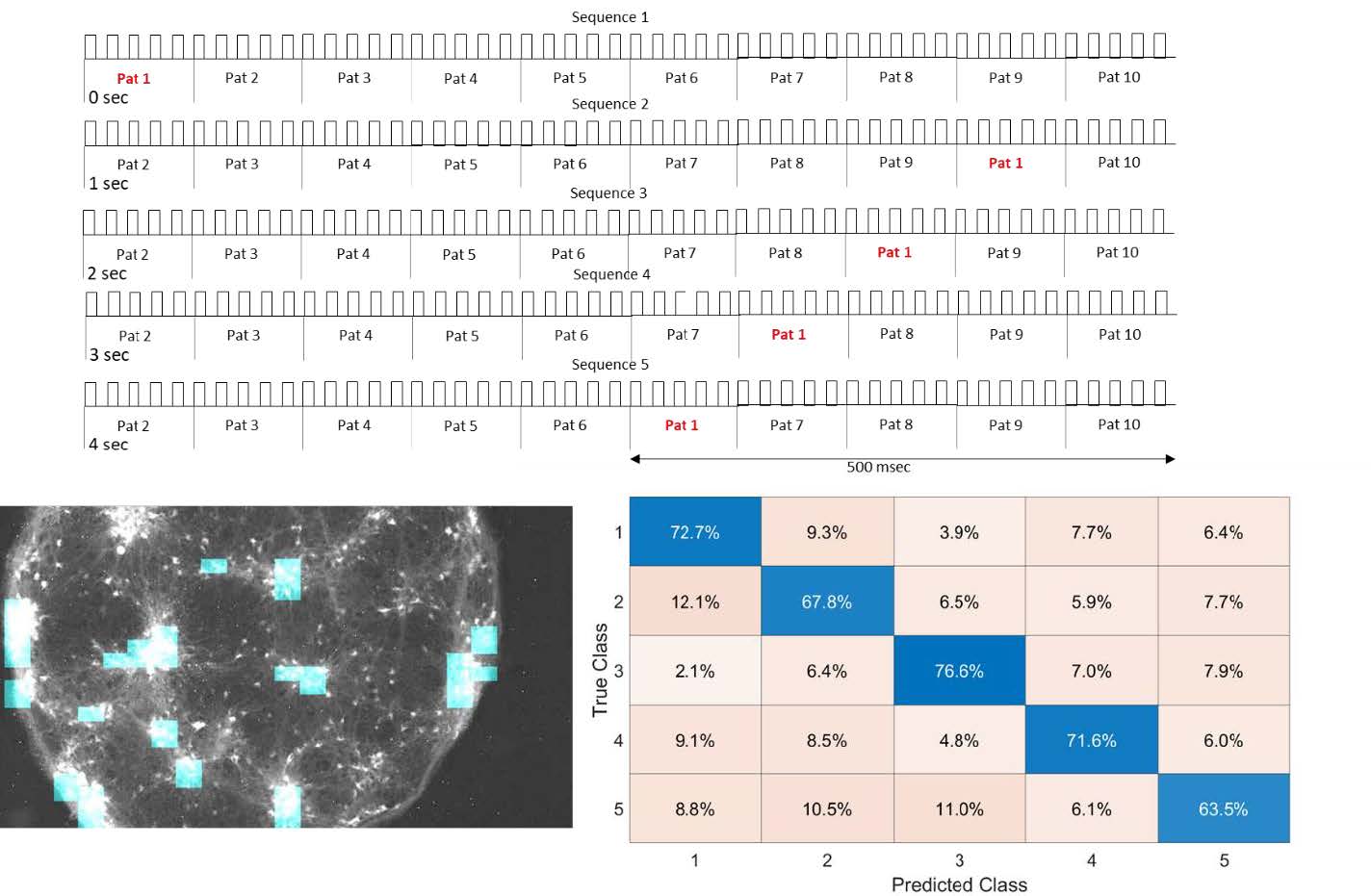
Neural Computation in Cortical Cultures
Confined neural networks in vitro with optical interface can be used to classify spatiotemporal information.
Patterned optogenetic stimulation to cultured ChR-expressing dissociated neurons resulted in suppression of population bursts. Strong correlation between outputs and input patterns that indicates processing of spatial information. Further analysis showed that the cortical network in vitro could classify and store information of spatiotemporal patterns up to 500 msec
By applying distributed optical stimulation patterns in the network, population bursts can be suppressed and simultaneously used to classify spatiotemporal information in the output neurons. These findings outline a paradigm of reservoir computing with living neural network which can be used to study spatiotemporal information in neural networks.
Courtesy Zubayer Ibne Ferdous & Dr. Yvgeny Berdichevsky, LeHigh University, USA
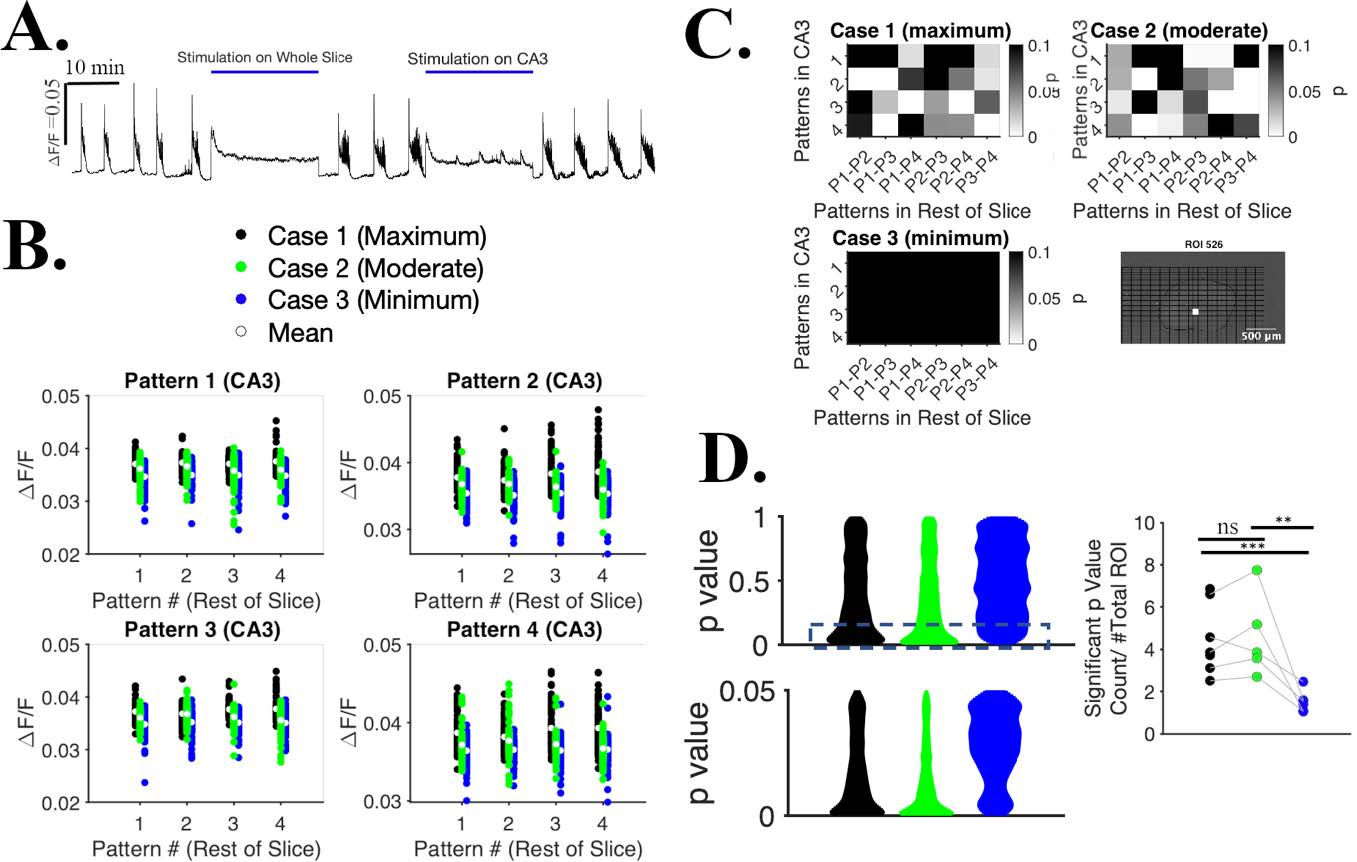
In-vitro Seizure Suppression by Patterned Optogenetic Stimulation while Allowing Information Processing in Hippocampus
Development of an effective method for suppressing seizures in such a way that activity in stimulated region is not completely shut off during the intervention. This allows stimulated area to participate in information processing and exchange of information with other brain regions, and may be a more gentle method of suppressing seizures than existing protocols.
Selective patterned optogenetic CA3 stimulation with ChR2 resulted reduction of seizure rate is lower and duration while not tetanizing the cells during stimulation and retaining the ability process other inputs during stimulation.
Courtesy Md Joynal Abedin & Dr. Yvgeny Berdichevsky, LeHigh University, USA
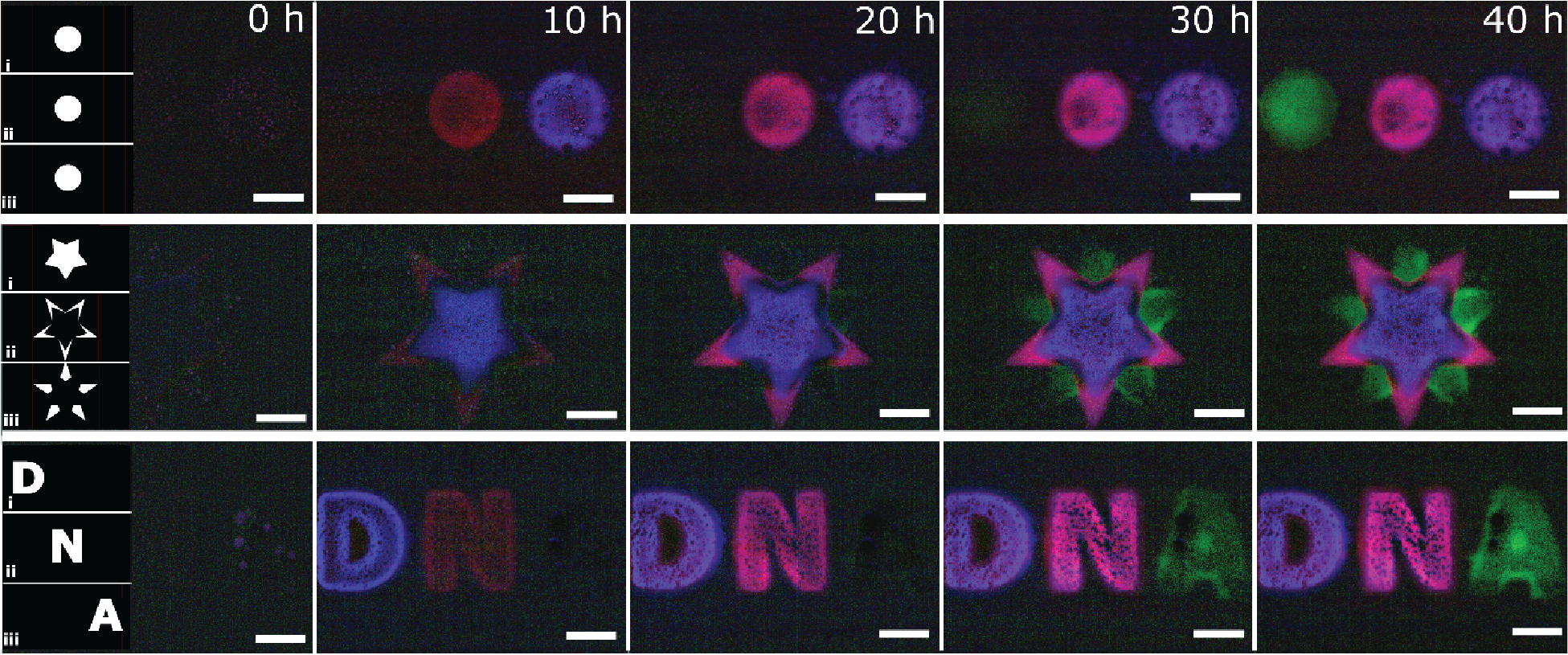
Photopatterning of Multi-Domain DNA Hydrogels
Fluorescent signals caused by the release of DNA from three 3-domain hydrogels that have unique embedded DNA within each domain.
Each row depicts the time-lapse for each multi-domain hydrogel, while each column represents the time of each photo. The blue domain activates first, followed by the red and then the green domain.
The digital masks uploaded to the DMD are shown in the left-most panel – each mask (row 1, 2, and 3) is used for the patterning of a hydrogel section with unique DNA molecules embedded within: Mask 1 contains red stage-1 DNA (released first), mask 2 contains blue stage-2 DNA (released second), and mask 3 contains green stage-3 DNA (released third). Scale bar is 200 µm.
Courtesy Misha Rubanov & Dr. Rebecca Schulman, Johns Hopkins University, USA

Photopatterning with mercaptopropylsilatrane and the Polygon 1000
By projecting different patterns, it is possible to control where molecules are attached, and so obtain monolayers of arbitrary shape. This results in monolayers with partial coverage – by breaking down a grayscale image into a series of different black and white images, researchers can individually control the exposure time at each pixel, and correspondingly tune the density of deposited dye molecules at each position.
The images are: 1) “Girl with a Pearl Earring” by Johannes Vermeer, 2) black-and-white version of “The Scream” by Edvard Munch, 3) “The Cardsharps” by Michelangelo Merisi da Caravaggio, and 4) the seal of Marquette University. Each image in the figure has an area of only 0.48 mm^2.
Courtesy Mark Mitmoen & Dr. Ofer Kedem, Marquette University, USA
Patterned Optogenetic Stimulation to Mimic Odors
The Mightex Polygon used to mimic “odors” by optogenetically activating spatio-temporal patterns in the early olfactory system of the mouse. Presentation of even monomolecular odorants already results in complex patterns of neural activity with unique but rigid spatiotemporal properties.
The Mightex Polygon provides full parametric control over both spatial and temporal properties of the optogenetic “odors” by precisely activating light spots over the dorsal olfactory bulb.
Courtesy Pedro Herrero-Vidal, Dr. Dima Rinberg, & Dr. Cristina Savin, New York University, USA
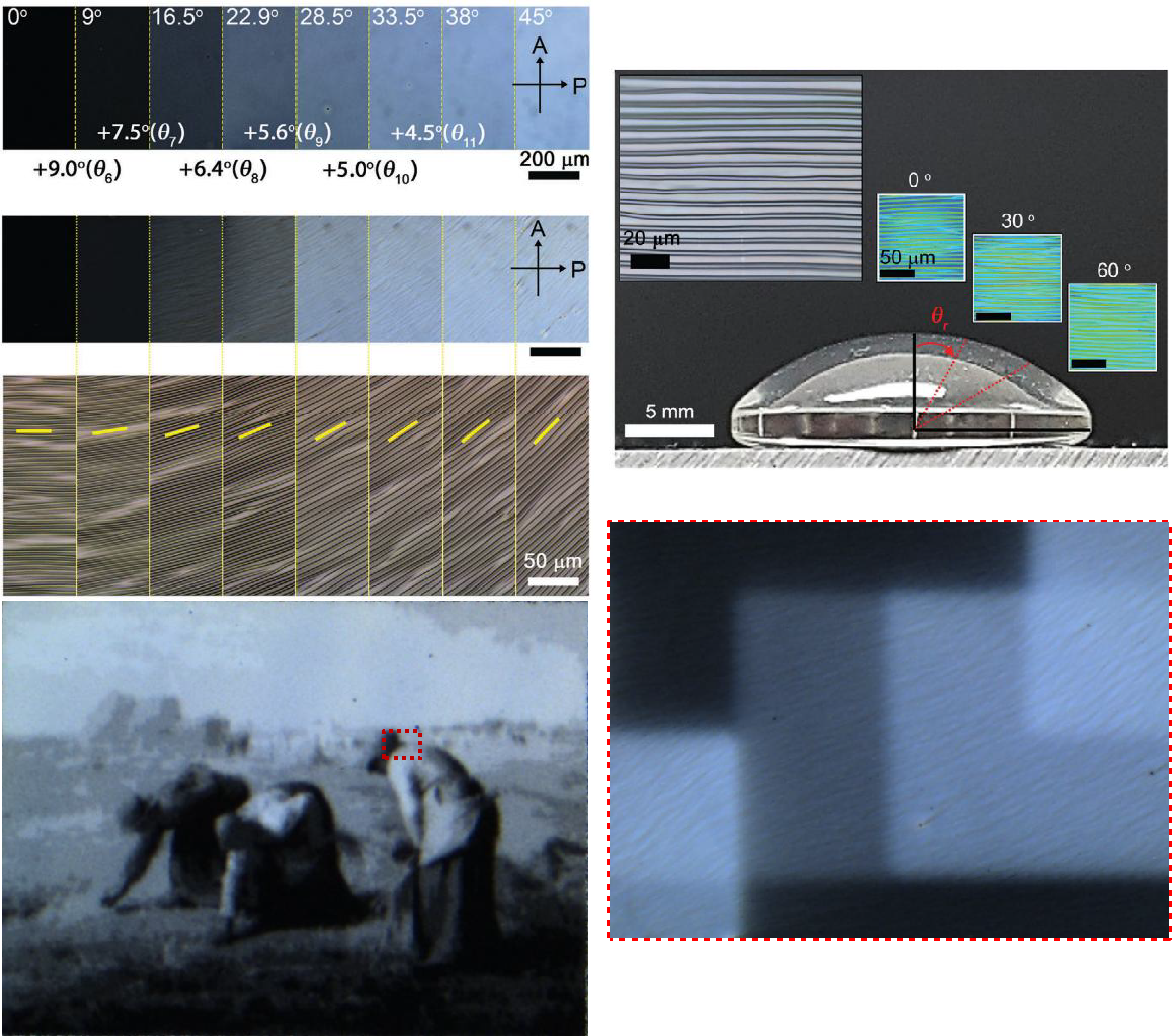

Pixelated Wrinkles Using Patterned Optical Phase Retardation
The Mightex’s Polygon1000 model used to form pixelated wrinkles on various surfaces. In addition, the continuous exposure technology of the Mightex Polygon has made it possible to locally expose the surface in conjunction with Nikon’s microscope software.
The optics based on a digital micro mirror technology (Polygon) assembles wrinkle pixels of a notably small dimension over a large area at fast fabrication speed. Furthermore, these pixelated wrinkles can be formed on curved geometries
Courtesy Kitae Kim & Dr. Jun-Hee Na, Chungnam National University, Korea
Patterned Optogenetics and Zebrafish Spinal Cord Organization
In vivo optogenetics and electrophysiology used to map connectivity of a major inhibitory population: V1 (En1+) neurons in larval zebrafish. Using the unique illumination patterns available with the Mightex Polygon, a 16×16 grid projected spanning approximately one spinal segment and sequentially activated 0-6 V1 neurons at a time. By translating this grid illumination to different spinal segments rostrally and caudally, connectivity tested from V1 neurons along significant lengths of the spinal cord.
Courtesy Dr. Mohini Sengupta & Dr. Martha Bagnall, WUSTL, USA
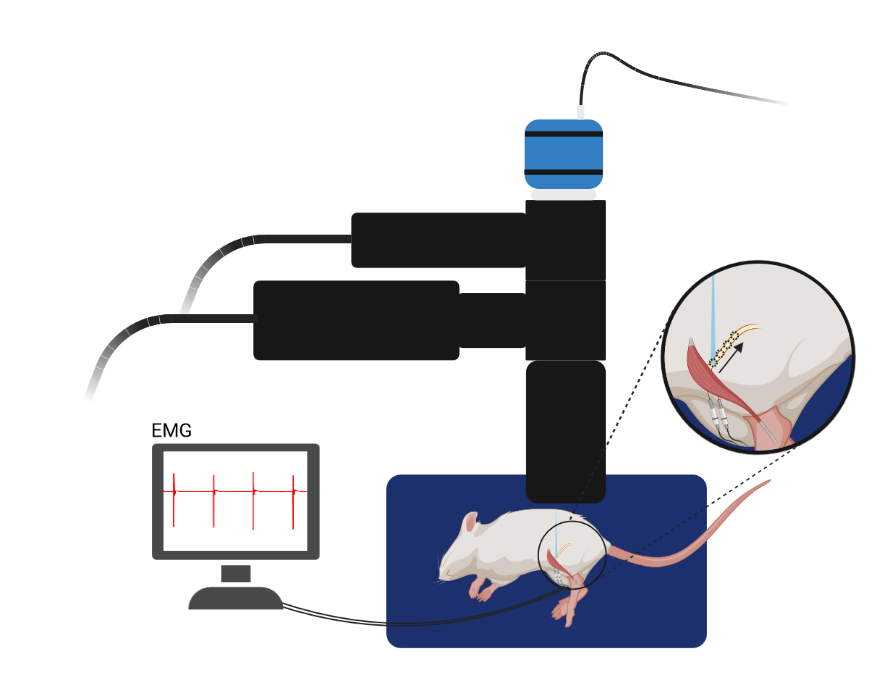
Patterned Optogenetics and Characterization of Muscular Responses
Polygon system used to stimulate the peroneal nerve in small segments starting from the distal muscle insertion moving proximally towards the sciatic nerve in a rat expressing Channelrhodopsin-2 (ChR2) in the peroneal nerve after an intramuscular virus injection in the Tibialis Anterior (TA) muscle. Simultaneous recording of muscle response to the optical stimulation in the TA muscle with intramuscular electromyography.
Courtesy Emma Moravec & Dr. Jordan Williams, Marquette University, USA
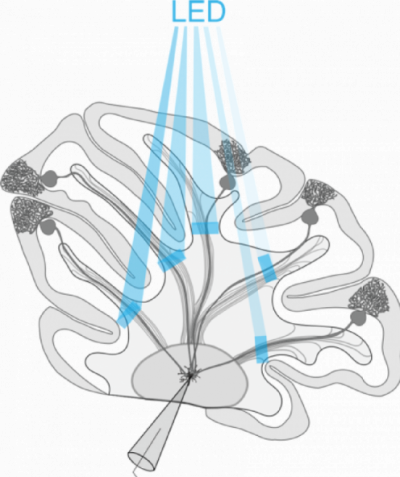
Patterned Optogenetics and Cerebellar Functional Connectivity
Patterned optogenetics stimulation of transgenically expressed ChR2 in Purkinje cells combined with electrophysiological recordings to evaluate patterns of connectivity between Purkinje cells and cerebellar nuclei neurons.
The Polygon is used to sequentially stimulate different cerebellar lobules in quick succession to assess the presence of a functional connection. These data are then used to characterize how different kinds of information are integrated within the cerebellum, and how these patterns may be implicated in behavior.
Courtesy Kim Gruver & Dr. Alanna Watt, McGill University, Canada
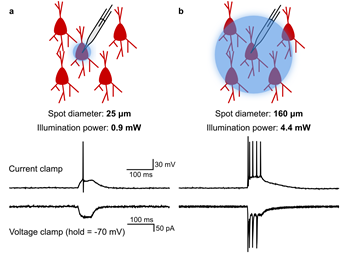
Single-Cell Resolution Optogenetic Spiking
(a) Illustration of highly targeted optical stimulation of a single ChR2-mCherry expressing neuron in the mouse somatosensory cortex. Using a small (25 μm diameter), low-powered (3 mW) spot of illumination centred on the target cell, action potentials could be induced in current clamp (top trace), without any indication in voltage clamp of post-synaptic currents caused by spiking in other neurons (bottom trace).
(b) Illustration of illumination with a large (125 μm diameter), high-powered (15 mW) spot. Multiple spikes were induced in current clamp, and voltage clamp traces showed evidence of post-synaptic currents caused by spiking in other neurons.
Courtesy Matthew Tran & Dr Blake Richards, University of Toronto, Canada.
ChR2 Assisted Circuit Mapping (CRACM)
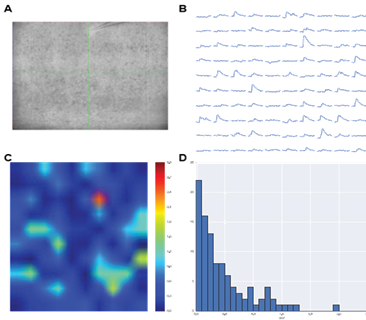
(A) Image of an acute brain slice prepared from a Thy1-ChR2-EYFP mouse with ChR2 expression in L5 pyramidal neurons. Whole-cell patch clamp recording from a L2/3 cell.
(B) Light-activated excitatory postsynaptic potentials (EPSPs) triggered by patterned illumination of a 10×10 grid with 473 nm LED.
(C) Colormap of the activation pattern.
(D) Histogram of automatically measured responses from all cells in a grid. Objective lens is 10X.
Courtesy Qiuyu Wu & Dr Alexander Chubykin, Purdue University, USA.
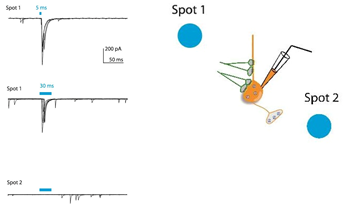
Optogenetic Stimulation of Synaptic Inputs
Light-evoked inhibitory postsynaptic current (IPSC) recorded in a transgenic mice expressing ChR2 in GABAergic neurons. The postsynaptic cell is non-GABAergic (ChR2 negative) and blue light stimulates GABAergic afferents expressing ChR2. Blue bars indicate the time of light illumination. Spot 1 illuminated by Polygon400 evoked reliable IPSCs whereas Spot 2 caused no response.
Courtesy Dr Wataru Inoue, University of Western Ontario, Canada.
Light-Modulated Protein-Protein Interactions on a Coverslip Surface
This video shows a protein-protein interaction that is inhibited by blue light exposure. One component is coupled to the coverslip through biotin surface chemistry, the other is labeled with mCherry, and binds to the coverslip in the dark. Patterned illumination with the Polygon400 results in revesible dissociation of the mCherry-tagged protein, which rebinds within minutes of turning off blue light exposure. The Wittmann lab is working on developing this into a cell adhesion surface that can be controlled by light.
Courtesy Dr Torsten Wittmann, University California San Francisco, USA.
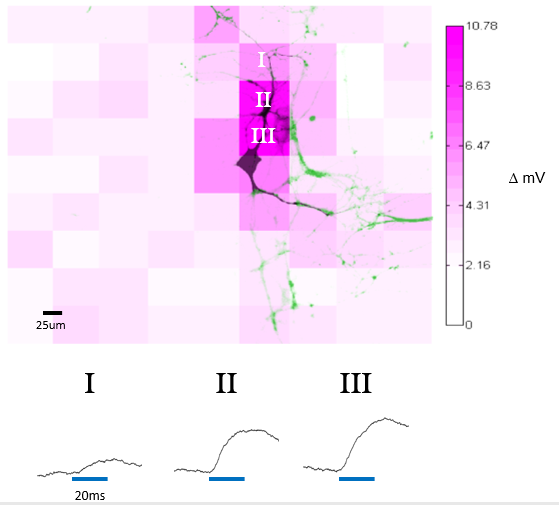
Mapping Optically Induced Depolarization in ChR2-expressing Hippocampal Neurons using Polygon400
E18 Sprague-Dawley rat neurons were transduced with CamKII-ChR2-GFP lentivirus. Somatic activity was recorded via whole-cell patch-clamp electrophysiology. Each field was illuminated by the Polygon400 at 100% power and 20ms exposure time. Intensity of magenta pattern represents depolarization with respect to the instantaneous resting potential prior to stimulation with the Polygon’s 470nm LED. In the image, green represents the magnitude of GFP signal and black represents the fluorescence intensity of AlexaFluor 594 backfilled by the patch pipette.
Courtesy Dr Jacob Robinson, Rice University, USA.
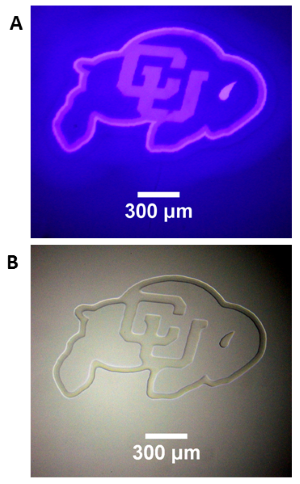
Photopatterning of an Acrylate Film
A 50 um film of liquid tetra (ethylen glycol) diacrylate, containg 1 wt% Irgacure 819 photointiator was irradiated through a glass coverslip, using the Polygon400 and a 400nm LED light source to project a pattern onto the film surface. (A) Projection of CU logo image using the 400nm LED and a 4X objective. (B) Brightfield image of the resulting pattern in the film. Photopolymerization causes a large change in refractive index in the resin, allowing immediate visualization of the pattern. Standard development techniques could subsequently be performed by washing the film in solvent to remove the unexposed areas of liquid resin.
Courtesy Gayla Berg, University of Colorado, USA.

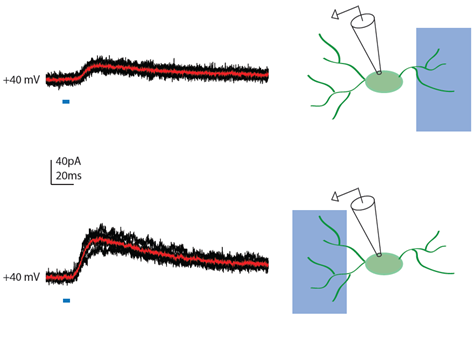
Achieving Subcellular Resolution of Input Activation in a 250um-thick Acute Cortical Section Using the Polygon400
NMDA-receptor mediated synaptic currents recorded at a +40mV membrane potential were elicited by light activation of channelrhodopsin-expressing terminals of thalamocortical afferents onto an upper layer cortical GABAergic interneuron. moving the 470nm rectangular illumination to the right or the left of the cell activates inputs of different strength innervating different somatodendritic domains of the cell. In black and red are the individual and the average traces respectively.
Courtesy Dr. Theofanis Karayannis, University of Zurich, Switzerland.
Publications by this Customer using Polygon:
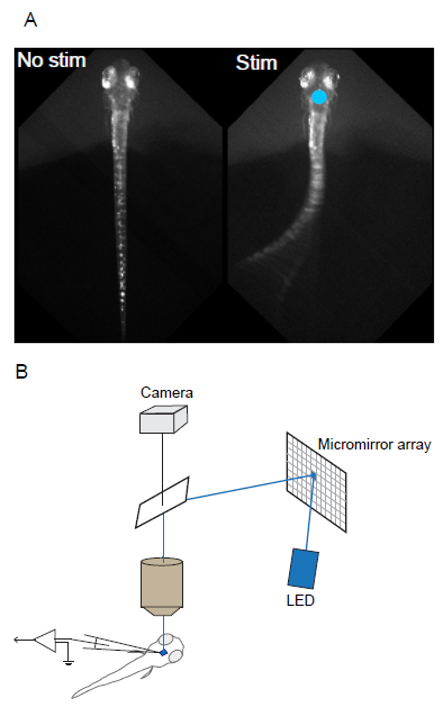
In Vivo Optogenetic Control of Zebrafish Larva Using Polygon400 Illuminator
A) Light-evoked response of a head-fixed larva expressing channelrhodopsin (right). Photostimulation site was indicated by a blue circle (470nm).
B) A schematic diagram of the Polygon400 patterned illumination in in vivo optogenetic mapping system.
Courtesy Dr. Sachiko Tsuda, Saitama University, Japan.
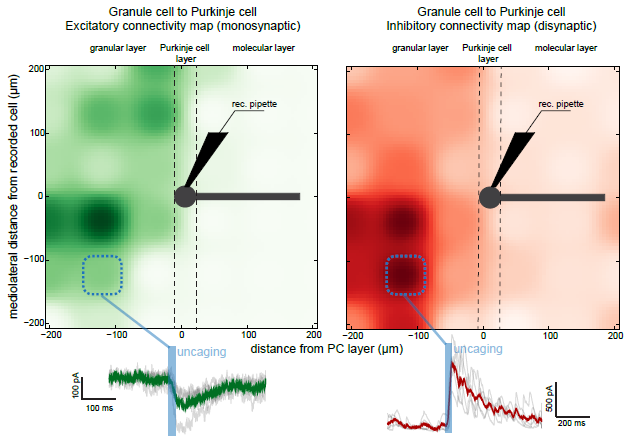
Uncaging RuBi Glutamate with the Polygon400 to Study Neuronal Networks in the Cerebellar Cortex
An example of connectivity mapping that allow to reproduce some results from in Valera et al. (elife, 2016). 100um RuBi glutamate was uncaged at various locations in the granular layer with 20ms pulses (blue bar), exciting notably granule cells. We can then measure the spatial organization of the granule cells to Purkinje cell (PC) connections by recording PCs in whole cell patch clamp. Granule cells triggers both monosynaptic excitatory current onto PCS (left map, measured a -60mW), but also disynaptic inhibitory currents via molecular layer interneurons (right map, measured at 0 mW). Average evoked responses at one location (dotted blue square) are shown at the bottom of the figure. Data storage, measurements, and map representations can be made online, using a homemade software in python.
Courtesy, Dr. Antoine Valera & Dr. Angus Silver, University College London, UK.
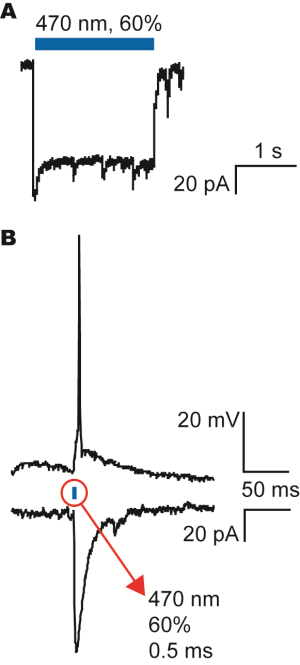
Optogenetic Activation of Channelrhodopsin in Transfected Hippocampal Neurons Using Mightex’s Polygon400
A) Patch-clamp recording of the current generated by the channelrhodopsin when activated with the blue LED light from the Polygon400 illuminator as indicated by the blue bar.
B) Activation of an action potential (upper trace) and the corresponding current under voltage-clamp conditions (lower trace). The action potential could be evoked by a 0.5ms stimulation of the blue LED light from the Polygon400 illuminator at 60% intensity.
Courtesy, Dr. Hans van Hooft, University of Amsterdam, Netherland.
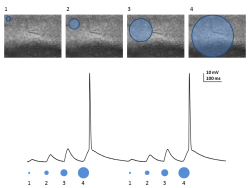
Using Polygon400 to Stimulate Hippocampal Slices Acutely Prepared from Mice Transgenically Expressing ChR2 in the Dentate Gyrus
The first figure a repetitive stimulation with a larger circular pattern which elicited increasing responses until an action potential is generated. The second figure is increasing the intensity of stimulation from 30% to 100% and keeping the size of the pattern the same. Here we’re driving the cell at 10Hz at 100% and one can see that the stimulation is sufficient to drive doublets of action potentials during each bout of depolorization.
Courtesy, Dr. Geoffrey G. Murphy, University of Michigan, USA.
Local Photostimulation of Channelrhodopsin-2 Using Polygon400 Illuminator
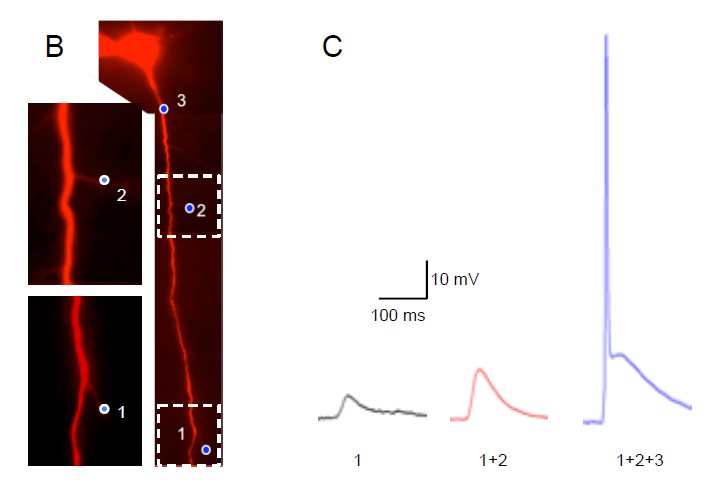
A) Acute brain slice from a YFP-channelrhodopsin-2 (ChR-2) mouse depicting its expression in cortical L5 pyramidal neurons. B) L5 pyramidal neuron from the somatosensory cortex filled with Alexa 594 to allow visualization of neuronal compartments without stimulating ChR-2. Blue dots indicate on the illuminated areas (Blue LED – 470 nm) along the apical dendrite. Expansion of the marked areas depicting the delicate dendrites that were stimulated. C) Electrophysiological (patch clamp) current recordings from the soma, corresponding to local photostimulation of ChR-2 by blue light. The numbers in the bottom of the trace corresponds to the stimulated dot numbers as indicated in B.
Courtesy, Dr. Yossi Buskila, University of Western Sydney, Australia.
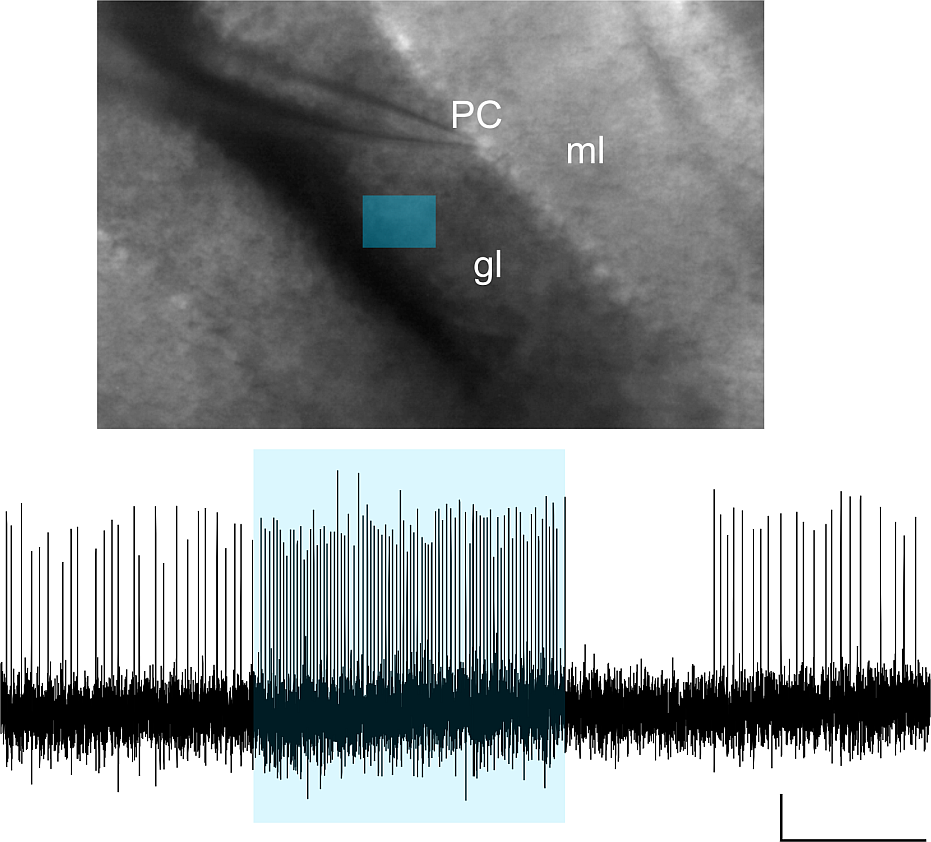
Purkinje Cell Firing Properties Unveiled by Optogenetic Activation of the Cerebellar Granular Layer Using Variable Light Patterns with Mightex Polygon400
Purkinje cell (PC) firing is monitored, while various light stimulation patterns are delivered to the granular layer (gl) and/or molecular layer (ml) of acute mouse cerebellar slices. The mice cerebellum was injected with AAVs carrying the ChR2 construct. The figure shows the firing frequency increase caused in a PC unit (scale bar 500ms) by optogenetic activation of a granular layer ROI (blue rectangle).
Courtesy, Dr. Lisa Mapelli & Dr. Simona Tritto, University of Pavia, Italy.
Relevant White Papers & Application Notes Using Polygon:
White Paper: Optical Patterned Illumination in Microfluidics
Publications
Polygon Related Publications
- Kim, K., & Na, J. H., Optical-Anisotropy-Exploited Irreproducible Functions Using Microwrinkles. ACS Applied Optical Materials, (2024).
- Zhao, Y., Huang, C. X., Gu, Y., Zhao, Y., Ren, W., Wang, Y., … & Song, J., Serotonergic modulation of vigilance states in zebrafish and mice. Nature Communications, 15(1), 2596, (2024).
- Zheng, J., Zhang, X. M., Tang, W., Li, Y., Wang, P., Jin, J., … & Li, B., An insular cortical circuit required for itch sensation and aversion. Current Biology, (2024).
- Li, X., You, J., Pan, Y., Song, C., Li, H., Ji, X., & Liang, F., Effective Regulation of Auditory Processing by Parvalbumin Interneurons in the Tail of the Striatum. Journal of Neuroscience, 44(5) (2024).
- Barua, N., Herken, A. M., Melendez-Velador, N., Platt, T. G., & Hansen, R. R. Photo-addressable microwell devices for rapid functional screening and isolation of pathogen inhibitors from bacterial strain libraries. Biomicrofluidics, 18(1), (2024).
- Wang, H., Wang, Q., Cui, L., Feng, X., Dong, P., Tan, L., … & Li, X. M., A molecularly defined amygdala-independent tetra-synaptic forebrain-to-hindbrain pathway for odor-driven innate fear and anxiety. Nature Neuroscience, 1-13, (2024).
- Matsumoto, A., Toyoshima, Y., Zhang, C., Isozaki, A., Goda, K., & Iino, Y., Neuronal sensorimotor integration guiding salt concentration navigation in Caenorhabditis elegans. Proceedings of the National Academy of Sciences, 121(5), e2310735121 (2024).
- Vu, M. A. T., Brown, E. H., Wen, M. J., Noggle, C. A., Zhang, Z., Monk, K. J., … & Howe, M. W., Targeted micro-fiber arrays for measuring and manipulating localized multi-scale neural dynamics over large, deep brain volumes during behavior. Neuron (2024).
- Wu, B., Rivas, D. P., & Das, S., Upstream mobility and swarming of light activated micromotors. Materials Advances (2024).
- Mulholland, H. N., Jayakumar, H., Farinella, D. M., & Smith, G. B., All-optical interrogation of millimeter-scale networks and application to developing ferret cortex. Journal of Neuroscience Methods, 403, 110051 (2023).
- Carlos A. Aguilar-Trigueros, Matthias C. Rillig, Jeff R., Powell Symbiotic status alters fungal eco-evolutionary offspring trajectories. Ecology Letters, 26, 9, 1483-1642 (2023).
- Naffaa, M. M., Khan, R. R., Kuo, C. T., & Yin, H. H., Cortical regulation of neurogenesis and cell proliferation in the ventral subventricular zone. Cell Reports, 42(7) (2023).
- Dema, A., Charafeddine, R., Rahgozar, S., van Haren, J., & Wittmann, T., Growth cone advance requires EB1 as revealed by genomic replacement with a light-sensitive variant. Elife, 12, e84143 (2023).
- Sheets, M. B., Tague, N., & Dunlop, M. J., An optogenetic toolkit for light-inducible antibiotic resistance. Nature Communications, 14(1), 1034 (2023).
- Chen, X., Chen, X., Elsayed, M., Edwards, H., Liu, J., Peng, Y., … & Wheeler, A. R., Steering Micromotors via Reprogrammable Optoelectronic Paths. ACS Nano, 17(6), 5894-5904 (2023).
- Tamatani, C., Shimba, K., Kotani, K., & Jimbo, Y., Activity-and Spatial-dependent Variations in Axonal Conduction Recorded from Microtunnel Electrodes. In 2023 15th Biomedical Engineering International Conference (BMEiCON) (pp. 1-3). IEEE, (2023).
- Xie, X., Hu, F., Zhou, Y., Liu, Z., Shen, X., Fu, J., … & Yu, Z. (2023). Photoswitchable Oxidopyrylium Ylide for Photoclick Reaction with High Spatiotemporal Precision: A Dynamic Switching Strategy to Compensate for Molecular Diffusion. Angewandte Chemie, 135(17), e202300034 (2023).
- Rivas, D. P., Sokolich, M., & Das, S., Spatial patterning of micromotor aggregation and flux. ChemNanoMat, 9(8), e202300225 (2023).
- McGowan, S. E., Discoidin domain receptor-2 enhances secondary alveolar septation in mice by activating integrins and modifying focal adhesions. American Journal of Physiology-Lung Cellular and Molecular Physiology, 324(3), L307-L324 (2023).
- Zhong, W., Zheng, W., & Ji, X., Spatial Distribution of Inhibitory Innervations of Excitatory Pyramidal Cells by Major Interneuron Subtypes in the Auditory Cortex. Bioengineering, 10(5), 547 (2023).
- Riback, J. A., Eeftens, J. M., Lee, D. S., Quinodoz, S. A., Donlic, A., Orlovsky, N., … & Brangwynne, C. P., Viscoelasticity and advective flow of RNA underlies nucleolar form and function. Molecular Cell, 83(17), 3095-3107 (2023).
- Lu, W., Lakonishok, M., & Gelfand, V. I., The dynamic duo of microtubule polymerase Mini spindles/XMAP215 and cytoplasmic dynein is essential for maintaining Drosophila oocyte fate. Proceedings of the National Academy of Sciences, 120(39), e2303376120 (2023).
- Mitmoen, M., & Kedem, O., UV-and Visible-Light Photopatterning of Molecular Gradients Using the Thiol–yne Click Reaction. ACS Applied Materials & Interfaces, 14(28), 32696-32705 (2022).
- Ji, X., Liu, W., Xiao, H., & Xiao, Z., The activated synaptic terminals beyond the light illumination range affect the results of optogenetics. NeuroReport, 33(7), 281-290 (2022).
- Wang, T., Yang, D., Chen, J., Chow, J., Hu, Y., Hoang, K., & Ansari, A., A Tetherless Microdriller for Maneuverability and On-Board Cargo Delivery Inside Viscoelastic Media. In 2022 International Conference on Manipulation, Automation and Robotics at Small Scales (MARSS) (pp. 1-6). IEEE (2022).
- Too, L. K., Shen, W., Protti, D. A., Sawatari, A., A. Black, D., Leamey, C. A., … & Simunovic, M. P., Optogenetic restoration of high sensitivity vision with bReaChES, a red-shifted channelrhodopsin. Scientific Reports, 12(1), 19312 (2022).
- Kumar, S., & Khammash, M., Platforms for optogenetic stimulation and feedback control. Frontiers in Bioengineering and Biotechnology, 10, 918917 (2022).
- Martinetti, L. E., Bonekamp, K. E., Autio, D. M., Kim, H. H., & Crandall, S. R., Short-term facilitation of long-range corticocortical synapses revealed by selective optical stimulation. Cerebral Cortex, 32(9), 1932-1949 (2022).
- Dema, A., van Haren, J., & Wittmann, T., Optogenetic EB1 inactivation shortens metaphase spindles by disrupting cortical force-producing interactions with astral microtubules. Current Biology, 32(5), 1197-1205 (2022).
- Gruber, A., Edri, O., Glatstein, S., Goldfracht, I., Huber, I., Arbel, G., … & Gepstein, L., Optogenetic Control of Human Induced Pluripotent Stem Cell‐Derived Cardiac Tissue Models. Journal of the American Heart Association, 11(4), e021615 (2022).
- Galloni, A. R., Ye, Z., & Rancz, E., Dendritic domain-specific sampling of long-range axons shapes feedforward and feedback connectivity of L5 neurons. Journal of Neuroscience, 42(16), 3394-3405 (2022).
- Liu, J., Enloe, C., Li-Oakey, K. D., & Oakey, J., Optimizing immunofunctionalization and cell capture on micromolded hydrogels via controlled oxygen-inhibited photopolymerization. ACS Applied Bio Materials, 5(10), 5004-5013 (2022).
- Shimba, K., Asahina, T., Sakai, K., Kotani, K., & Jimbo, Y., Recording saltatory conduction along sensory axons using a high-density microelectrode array. Frontiers in Neuroscience, 16, 854637 (2022).
- Liu, Y., Li, S., Zhang, X., Wang, L., Li, Z., Wu, W., … & Yang, Y., Corticotropin releasing factor neurons in the visual cortex mediate long-term changes in visual function induced by early adversity. Neurobiology of Stress, 21, 100504 (2022).
- Kishi, J. Y., Liu, N., West, E. R., Sheng, K., Jordanides, J. J., Serrata, M., … & Yin, P., Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing. Nature Methods, 19(11), 1393-1402 (2022).
- Zhang, S., Elsayed, M., Peng, R., Chen, Y., Zhang, Y., Neale, S. L., & Wheeler, A. R., Influence of light pattern thickness on the manipulation of dielectric microparticles by optoelectronic tweezers. Photonics Research, 10(2), 550-556 (2022).
- Jack, A., Kim, Y., Strom, A. R., Lee, D. S., Williams, B., Schaub, J. M., … & Brangwynne, C. P., Compartmentalization of telomeres through DNA-scaffolded phase separation. Developmental Cell, 57(2), 277-290 (2022).
- Honda, M., Kimura, R., Harada, A., Maehara, K., Tanaka, K., Ohkawa, Y., & Oki, S., Photo-isolation chemistry for high-resolution and deep spatial transcriptome with mouse tissue sections. STAR protocols, 3(2), 101346 (2022).
- Sengupta, M., Daliparthi, V., Roussel, Y., Bui, T. V., & Bagnall, M. W., Spinal V1 neurons inhibit motor targets locally and sensory targets distally. Current Biology, 31(17), 3820-3833 (2021).
- Gruber, A., Edri, O., Huber, I., Arbel, G., Gepstein, A., Shiti, A., … & Gepstein, L., Optogenetic modulation of cardiac action potential properties may prevent arrhythmogenesis in short and long QT syndromes. JCI Insight, 6(11) (2021).
- Honda, M., Oki, S., Kimura, R., Harada, A., Maehara, K., Tanaka, K., … & Ohkawa, Y., High-depth spatial transcriptome analysis by photo-isolation chemistry. Nature Communications, 12(1), 4416 (2021).
- Dine, E., Reed, E. H., & Toettcher, J. E., Positive feedback between the T cell kinase Zap70 and its substrate LAT acts as a clustering-dependent signaling switch. Cell Reports, 35(12) (2021).
- Lemaire, L., Desroches, M., Krupa, M., Pizzamiglio, L., Scalmani, P., & Mantegazza, M., Modeling NaV1. 1/SCN1A sodium channel mutations in a microcircuit with realistic ion concentration dynamics suggests differential GABAergic mechanisms leading to hyperexcitability in epilepsy and hemiplegic migraine. PLoS Computational Biology, 17(7), e1009239 (2021).
- Chua, C. J., Han, J. L., Li, W., Liu, W., & Entcheva, E., Integration of engineered “spark-cell” spheroids for optical pacing of cardiac tissue. Frontiers in Bioengineering and Biotechnology, 9, 658594 (2021).
- Jiang, H. H., Guo, A., Chiu, A., Li, H., Lai, C. S. W., & Lau, C. G., Target-specific control of piriform cortical output via distinct inhibitory circuits. FASEB Journal, 35(10), e21944 (2021).
- McFann, S., Dutta, S., Toettcher, J. E., & Shvartsman, S. Y., Temporal integration of inductive cues on the way to gastrulation. Proceedings of the National Academy of Sciences, 118(23), e2102691118 (2021).
- Zhang, S., Li, W., Elsayed, M., Peng, J., Chen, Y., Zhang, Y., … & Wheeler, A. R., Integrated assembly and Photopreservation of topographical micropatterns. Small, 17(37), 2103702 (2021).
- Zhang, S., Elsayed, M., Peng, R., Chen, Y., Zhang, Y., Peng, J., … & Wheeler, A. R., Reconfigurable multi-component micromachines driven by optoelectronic tweezers. Nature Communications, 12(1), 5349 (2021).
- Bellot-Saez, A., Stevenson, R., Kékesi, O., Samokhina, E., Ben-Abu, Y., Morley, J. W., & Buskila, Y., Neuromodulation of astrocytic K+ clearance. International Journal of Molecular Sciences, 22(5), 2520 (2021).
- Chenais N.A.L., Leccardi M.J.I.A., Ghezzi D. Photovoltaic Retinal Prosthesis Restores High-Resolution Responses to Single-Pixel Stimulation in Blind Retinas. (2021) Communications Materials.
- Ruan H., Wang L., Yuan F., Weng S., Zhong Y. Orexin-A Differentially Modulates Inhibitory and Excitatory Synaptic Transmission in Rat Inner Retina. (2021) Neuropharmacology.
- Lee D.S.W, Wingreen N.S., Brangwynne C.P. Chromatin Mechanics Dicates Subdiffusion and Coarsening Dynamics of Embedded Condensates. (2021). Nature Physics.
- Lu M., van Tartwijk F.W., Lin J.Q., Nijenhuis W., Kaminski C.F. The Structure and Global Distribution of the Endoplasmic Reticulum Network are Actively Regulated by Lysosomes. (2021) Science Advances.
- Geisterfer Z.M., Oakley J., Gatlin J.C. Microfluidic Encapsulation of Xenopus laevis Cell-Free Extracts Using Hydrogel Photolithography. (2020) STAR Protocols.
- Tabarean I.V. Activation of Preoptic Arginine Vasopressin Neurons Induces Hyperthermia in Male Mice. (2020) Endocrinology.
- Chenais N.A.L., Leccardi M.J.I.A., Ghezzi D. Naturalistic Spatiotemporal Modulation of Epiretinal Stimulation Increases the Response Persistence of Retinal Ganglion Cell. (2020) Journal of Neural Engineering.
- Wei L., Li W., Entcheva E., Li Z. Microfluidics-Enabled 96-Well Perfusion System for High-Throughput Tissue Engineering and Long-Term All-Optical Electrophysiology. (2020) Lab on a Chip.
- Kastli R., Vighagen R., van der Bourg A., Argunsah A.O.,Aguzzi A., Helmchen F., Karayannis T. Developmental Divergence of Sensory Stimulus Representation in Cortical Interneurons. (2020) Nature Communications.
- Anastasiades P.G., Collins D.P., Carter A.G. Mediodorsal and Ventromedial Thalamus Engage Distinct L1 Circuits in the Prefrontal Cortex. (2020) Neuron.
- Liu X., Dimidschstein J., Fishell G., Carter A.G. Hippocampal Inputs Engage CCK+ Interneurons to Mediate Endocannabinoid-Modulated Feed-Forward Inhibition in the Prefrontal Cortex. (2020) eLife.Optogenetic Sensor: A Glutamine Lever Induces Unfolding of the Jα Helix. (2020) ACS Chemical Biology.
- Fattahi N., Nieves-Otero P.A., Masigol M., van der Vlies A.J., Jensen R.S., Hansen R.R., Platt T.G. Photodegradable Hydrogels for Rapid Screening, Isolation, and Genetic Characterization of Bacteria with Rare Phenotypes. (2020) BioMacromolecules.
- Lindner M., Gilhooley M.J., Peirson S.N., Hughes S., Hankins M.W. The Functional Characteristics of Optogenetic Gene Therapy for Vision Restoration. (2020)Cellular and Molecular Life Sciences.
- Johnson H.E., Djabrayan N.J.V., Shvartsman S.Y., Toettcher J.E. Optogenetic Rescue of a Patterning Mutant. (2020) Current Biology.
- Ravindran P.T., Wilson M.Z., Jena S.G., Toettcher J.E. Engineering Combinatorial and Dynamic Decoders Using Synthetic Immediate-Early Genes. (2020)Communications Biology.
- Carrasco-Lopez C., Alam N., Toettcher J.E., Avalos J.L. Development of Light-Responsive Protein Binding in the Monobody Non-Immunoglobulin Scaffold. (2020) Nature Communications.
- Murphy C.A., Raz A., Grady S.M., Banks M.I. Optogenetic Activation of Afferent Pathways in Brain Slices and Modulation of Responses by Volatile Anesthetics. (2020) JOVE.
- Cheng C., Harpster M.H., Oakey J. Convection-Driven Microfabricated Hydrogels for Rapid Biosensing. (2020) Analyst.
- Sulerud T., Sami A.B., Li G., Kloxin A., Oakey J., & Gatlin J. Microtubule-Dependent Pushing Forces Contribute to Long-Distance Aster Movement and Centration in Xenopus Laevis Egg Extracts. (2020) Molecular Biology of the Cell.
- Iuliano J.N., Collado J.T., Gil A.A., Ravindran P.T.,Gardner K.H., Simmerling C.L., Meech S.R., Tonge P.J. Unraveling the Mechanism of a LOV Domain Optogenetic Sensor: A Glutamine Lever Induces Unfolding of the Jα Helix. (2020) ACS Publications.
- Wang L., Guo W., Shen X.,Lyu Q., Herbison A.E., Kuang Y. Different Dendritic Domains of the GnRH Neuron Underlie the Pulse and Surge Modes of GnRH Secretion in Female Mice. (2020) eLife.
- Liu S. Dopamine Suppresses Synaptic Responses of Fan Cells in the Lateral Entorhinal Cortex to Olfactory Bulb Input in Mice. (2020) Frontiers in Cellular Neuroscience.
- Chong E., Moroni M., Wilson C., Shoham S., Panzeri S., Rinberg D. Manipulating Synthetic Optogenetic Odors Reveals the Coding Logic of Olfactory Perception.(2020) Science.
- Geisterfer Z.M., Zhu D.Y., Mitchison T.J., Oakey J., Gatlin J.C. Microtubule Growth Rates Are Sensitive to Global and Local Changes in Microtubule Plus-End Density. (2020) Current Biology.
- Kauvar I.V., Machado T.A., Yuen E., Kochalka J., Choi M., Allen W.E., Wetzstein G., Deisseroth K. Cortical Observation by Synchronous Multifocal Optical Sampling Reveals Widespread Population Encoding of Actions. (2020) Neuron.
- Mu M., Geng H., Rong K., Peng R., Wang S., Geng L., Qian Z., Yung W., Ke Ya. A Limbic Circuitry Involved in Emotional Stress-Induced Grooming. (2020) Nature Communications
- Song Y., Hwang Y., Kim K., Lee H.,Jung M.W., Petersen C.C.H., Knott G., Lee S., Lee S. Somatostatin Enhances Visual Processing and Perception by Suppresing Excitatory Inputs to Parvalbumin-positive Interneurons in V1. (2020) Science Advances
- Kissinger S.T., Anderson A.K., Pak A., & Chubykin A.A. Visual Experience-Dependent Oscillations and Underlying Circuit Connectivity Changes Are Impaired in Fmr1 KO Mice. (2020) Cell Reports
- Oboti L. & Sokolowski K. Gradual Wiring of Olfactory Input to Amygdala Feedback Circuits. (2020) Scientific Reports
- Goglia A.G., Wilson M.Z., Sibert J., Basta L.P., Deveport D., & Toettcher J.E. A Live-Cell Screen for Altered Erk Dynamics Reveals Principles of Proliferative Control. (2020) Cell Systems
- Jiang Z., McBride R., Jiang K., Oakey J. Crosslinker Length Dictates Step-Growth Hydrogel Network Formation Dynamics and Allows Rapid On-Chip Photoencapsulation. (2020) Biofabrication
- Chung H., Park K., Jang H.J., Kohl M.M., & Kwag J. Dissociation of Somatostatin and Parvalbumin Interneurons Circuit Dysfunctions Underlying Hippocampal Theta and Gamma Oscillations Impaired by Amyloid β Oligomers In Vivo. (2020) Brain Structure and Function
- Nijenhuis W., Grinvsven M.M.P., & Kapitein L.C. An Optimized Toolbox for the Optogenetic Control of Intracellular Transport. (2020) Journal of Cell Biology
- Papasavvas C.A., Parrish R., & Trevelyan A.J. Propagating Activity in Neocortex, Mediated by Gap-Junctions and Modulated by Extracellular Potassium. (2020) eNeuro
- Callahan R.A., Roberts R., Sengupta M., Kimura Y., Higashijima S. & Bagnall M.W. Spinal V2b Neurons Reveal a Role for Ipsilateral Inhibition in Speed Control. (2020) eLife
- Murphy C., Krause B., & Banks M. Selective Effects of Isoflurane on Cortico-Cortical Feedback Afferent Responses in Murine Non-Primary Neocortex. (2019) British Journal of Anaesthesia.
- Dorsey P.J., Rubanov M., Wang W., & Schulman R. Digital Maskless Photolithographic Patterning of DNA-Functionalized Poly(ethylene glycol) Diacrylate Hydrogels with Visible Light Enabling Photodirected Release of Oligonucleotides. (2019) ACS Macro Letters.
- Petrovskaya L.E., Roshchin M.V., Smirnova G.R., Kolotova D.E., Balaban P.M., Ostrovsky M.A., & Malyshev A.Y. Bicistronic Construct for Optogenetic Prosthesis of Ganglion Cell Receptive Field of Degenerative Retina. (2019) Doklady Biochemistry and Biophysics.
- Yoon J.Y., Lee H.R., Ho W.K., & Lee S.H. Disparities in Short-Term Depression Among Prefrontal Cortex Synapses Sustain Persistent Activity in a Balanced Network. (2019) Cerebral Cortex.
- Noren B.E., Shaha R.K., Stenquist A.T., Frick C.P., & Oakey J.S. Cell Printing in Complex Hydrogel Scaffolds. (2019) IEEE Transactions of Nanobioscience.
- Wei Q., Krolewski D.M., Moore S., Kumar V., Li F., Martin B., Tomer R., Murphy G.G., Deisseroth K., Watson Jr S.J., & Akil H. Uneven Balance of Power Between Hypothalamic Peptidergic Neurons in the Control of Feeding. (2018) Proceedings of the National Academy of Sciences.
- Moscato L., Montagna I., De Propris L., Tritto S., Mapelli L., & D’Angelo E. Long-Lasting Response Changes in Deep Cerebellar Nuclei in vivo Correlate with Low-Frequency Oscillations. (2019) Frontiers in Cellular Neuroscience.
- Johnson H.E. & Toettcher J.E. Signaling Dynamics Control Cell Fate in the Early Drosophila Embryo. (2019) Developmental Cell.
- Simonova N.A., Bal N.V., Balaban P.M., Volgushev M.A., & Malyshev A.Y. An Optogenetic Approach to Studies of the Mechanisms of Heterosynaptic Plasticity in Neocortical Neurons. (2019) Neuroscience and Behavioral Physiology.
- Shen C., Zheng D., Li K., Yang J., Pan H., Yu X., Fu J., Zhu Y., Sun Q., Tang M., Zhang Y., Sun P., Xie Y., Duan S., Hu H., & Li X. Cannabinoid CB1 Receptors in the Amygdalar Cholecystokinin Glutamatergic Afferents to Nucleus Accumbens Modulate Depressive-Like Behavior. (2019) Nature Medicine.
- Li Y., Li C., Xi W., Jin S., Wu Z., Jiang P., Dong P., He X., Xu F., Duan S., Zhou Y., & Li X. Rostral and Caudal Ventral Tegmental Area GABAergic Inputs to Different Dorsal Raphe Neurons Participate in Opioid Dependence. (2019) Neuron.
- van Der Vlies J.A., Barua, N., Nieves-Otero P.A., Plat T.G., & Hansen R.R. On Demand Release and Retrieval of Bacteria from Microwell Arrays Using Photodegradable Hydrogel Membranes. (2018) ACS Applied Bio Materials.
- Ogasawara, S. Transcription Driven by Reversible Photocontrol of Hyperstable G-Quadruplexes. (2018) ACS Synthetic Biology.
- Konetski D., Zhang D., Schwartz D.K., & Bowman C.N. Photo-induced Pinocytosis for Artificial and Proto-Cell Systems. (2018) Chemistry of Materials.
- Majumder R., Feola I., Teplenin A.S., de Vries A.A., Panfilov A.V., & Pijnappels D. Optogenetics Enables Real-Time Spatiotemporal Control Over Spiral Wave Dynamics in an Excitable Cardiac System. (2018) eLife.
- Tabor, K.M., Smith, T.S., Brown, M., Bergeron, S.A., Briggman, K.L., & Burgess H.A. Presynaptic Inhibition Selectively Gates Auditory Transmission to the Brainstem Startle Circuit. (2018) Current Biology.
- Virag, T.T., Cserep, C., Schlingloff, D., Posfai, B., Szonyi, A., Sos, K.E., Kornyei, Z., Denes, A., Gulyas, A.I., Freund, T.F., & Nyiri, G. Co-Transmission of Acetlycholine and GABA Regulates Hippocampal States. (2018) Nature Communications.
- Rhomberg, T., Rovira-Esteban, L., Vikor, A., Paradiso, E., Kremser C., Nagy-Pal, P., Papp, O.I., Tasan, R., Erdelyi, F., Szabo, G., Ferraguit, F. & Hajor, N. VIP-Immunoreactive Interneurons Within Circuits of the Mouse Basolateral Amygdala. (2018) Journal of Neuroscience.
- Teplenin, A.S., Dierckx, H., de Vries, A.A.F., Pijnappels, D.A., & Panfilov, A.V. Paradoxical Onset of Arrhythimic Waves from Depolarizes Areas in Cardiac Tissue Due to Curvature-Dependent Instability. (2018) Physical Review X.
- Che, A., Babij, R., Iannone, A.F., Fetcho, R.N., Ferrer, M., Liston, C., Fishell, G., & De Marco Garcia, N.V. Layer I Interneurons Sharpen Sensory Maps during Neonatal Development. (2018) Neuron.
- Johnson H.E., Djabrayan N.J.V., Shvartsman S.Y., Toettcher J.E. Optogenetic Rescue of a Patterning Mutant. (2020) Current Biology.
- Fattahi N., Nieves-Otero P.A., Masigol M.,Jensen R.S., Hansen R.R., Platt T.G. Photodegradable Hydrogels for Rapid Screening, Isolation, and Genetic Characterization of Bacteria with Rare Phenotypes. (2020) BioMacromolecules.
- Wang L., Guo W., Shen X., Yeo S., Long H., Wang Z., Lyu Q., Herbison A.E., Kuang Y. Different Dendritic Domains of the GnRH Neuron Underlie the Pulse and Surge Modes of GnRH Secretion in Female Mice. (2020) eLife.
- Liu S. Dopamine Suppresses Synaptic Responses of Fan Cells in the Lateral Entorhinal Cortex to Olfactory Bulb Input in Mice. (2020) Frontiers in Cellular Neuroscience.
- Chong E., Moroni M., Wilson C., Shoham S., Panzeri S., Rinberg D. Manipulating Synthetic Optogenetic Odors Reveals the Coding Logic of Olfactory Perception. (2020) Science.
- Geisterfer Z.M., Zhu D.Y., Mitchison T.J., Oakey J., Gatlin J.C. Microtubule Growth Rates Are Sensitive to Global and Local Changes in Microtubule Plus-End Density. (2020) Current Biology.
- Kauvar I.V., Machado T.A., Yuen E., Kochalka J., Choi M., Allen W.E., Wetzstein G., Deisseroth K. Cortical Observation by Synchronous Multifocal Optical Sampling Reveals Widespread Population Encoding of Actions. (2020) Neuron.
- Mu M., Geng H., Rong K., Peng R., Wang S., Geng L., Qian Z., Yung W., Ke Ya. A Limbic Circuitry Involved in Emotional Stress-Induced Grooming. (2020) Nature Communications
- Song Y., Hwang Y., Kim K., Lee H., Kim J., Maclachlan C., Dubois A., Jung M.W., Petersen C.C.H., Knott G., Lee S., Lee S. Somatostatin Enhances Visual Processing and Perception by Suppresing Excitatory Inputs to Parvalbumin-positive Interneurons in V1. (2020) Science Advances
- Kissinger S.T., Wu Q., Quinn C.J., Anderson A.K., Pak A., & Chubykin A.A. Visual Experience-Dependent Oscillations and Underlying Circuit Connectivity Changes Are Impaired in Fmr1 KO Mice. (2020) Cell Reports
- Oboti L. & Sokolowski K. Gradual Wiring of Olfactory Input to Amygdala Feedback Circuits. (2020) Scientific Reports
- Callahan R.A., Roberts R., Sengupta M., Kimura Y., Higashijima S. & Bagnall M.W. Spinal V2b Neurons Reveal a Role for Ipsilateral Inhibition in Speed Control. (2020) eLife
- Dorsey P.J., Rubanov M., Wang W., & Schulman R. Digital Maskless Photolithographic Patterning of DNA-Functionalized Poly(ethylene glycol) Diacrylate Hydrogels with Visible Light Enabling Photodirected Release of Oligonucleotides. (2019) ACS Macro Letters.
- Petrovskaya L.E., Roshchin M.V., Smirnova G.R., Kolotova D.E., Balaban P.M., Ostrovsky M.A., & Malyshev A.Y. Bicistronic Construct for Optogenetic Prosthesis of Ganglion Cell Receptive Field of Degenerative Retina. (2019) Doklady Biochemistry and Biophysics.
- Goglia A.G., Wilson M.Z., Sibert J., Basta L.P., Deveport D., & Toettcher J.E. A Live-Cell Screen for Altered Erk Dynamics Reveals Principles of Proliferative Control. (2020) Cell Systems
- Jiang Z., Shaha R., McBride R., Jiang K., Tang M., Xu B., Goroncy A.K., Frick C., & Oakey J. Crosslinker Length Dictates Step-Growth Hydrogel Network Formation Dynamics and Allows Rapid On-Chip Photoencapsulation. (2020) Biofabrication
- Chung H., Park K., Jang H.J., Kohl M.M., & Kwag J. Dissociation of Somatostatin and Parvalbumin Interneurons Circuit Dysfunctions Underlying Hippocampal Theta and Gamma Oscillations Impaired by Amyloid β Oligomers In Vivo. (2020) Brain Structure and Function
Nijenhuis W., Grinvsven M.M.P., & Kapitein L.C. An Optimized Toolbox for the Optogenetic Control of Intracellular Transport. (2020) Journal of Cell Biology - Papasavvas C.A., Parrish R., & Trevelyan A.J. Propagating Activity in Neocortex, Mediated by Gap-Junctions and Modulated by Extracellular Potassium. (2020) eNeuro
- Yoon J.Y., Lee H.R., Ho W.K., & Lee S.H. Disparities in Short-Term Depression Among Prefrontal Cortex Synapses Sustain Persistent Activity in a Balanced Network. (2019) Cerebral Cortex.
- Noren B.E., Shaha R.K., Stenquist A.T., Frick C.P., & Oakey J.S. Cell Printing in Complex Hydrogel Scaffolds. (2019) IEEE Transactions of Nanobioscience.
- Moscato L., Montagna I., De Propris L., Tritto S., Mapelli L., & D’Angelo E. Long-Lasting Response Changes in Deep Cerebellar Nuclei in vivo Correlate with Low-Frequency Oscillations. (2019) Frontiers in Cellular Neuroscience.
- Murphy C., Krause B., & Banks M. Selective Effects of Isoflurane on Cortico-Cortical Feedback Afferent Responses in Murine Non-Primary Neocortex. (2019) British Journal of Anaesthesia.
- Johnson H.E. & Toettcher J.E. Signaling Dynamics Control Cell Fate in the Early Drosophila Embryo. (2019) Developmental Cell.
- Simonova N.A., Bal N.V., Balaban P.M., Volgushev M.A., & Malyshev A.Y. An Optogenetic Approach to Studies of the Mechanisms of Heterosynaptic Plasticity in Neocortical Neurons. (2019) Neuroscience and Behavioral Physiology.
- Shen C., Zheng D., Li K., Yang J., Pan H., Tang M., Zhang Y., Sun P., Xie Y., Duan S., Hu H., & Li X. Cannabinoid CB1 Receptors in the Amygdalar Cholecystokinin Glutamatergic Afferents to Nucleus Accumbens Modulate Depressive-Like Behavior. (2019) Nature Medicine.
- Li Y., Li C., Xi W., Jin S., Wu Z., Jiang P., Dong P., He X., Xu F., Duan S., Zhou Y., & Li X. Rostral and Caudal Ventral Tegmental Area GABAergic Inputs to Different Dorsal Raphe Neurons Participate in Opioid Dependence. (2019) Neuron.
- Mathejczyk, T. F., & Wernet, M. F., Heading choices of flying Drosophila under changing angles of polarized light. Scientific Reports, 9(1), 16773 (2019).
- van Der Vlies J.A., Barua, N., Nieves-Otero P.A., Plat T.G., & Hansen R.R. On Demand Release and Retrieval of Bacteria from Microwell Arrays Using Photodegradable Hydrogel Membranes. (2018) ACS Applied Bio Materials.
- Konetski D., Zhang D., Schwartz D.K., & Bowman C.N. Photo-induced Pinocytosis for Artificial and Proto-Cell Systems. (2018) Chemistry of Materials.
Ogasawara, S. Transcription Driven by Reversible Photocontrol of Hyperstable G-Quadruplexes. (2018) ACS Synthetic Biology. - Wei Q., Krolewski D.M., Moore S., Kumar V., Li F., Martin B., Tomer R., Murphy G.G., Deisseroth K., Watson Jr S.J., & Akil H. Uneven Balance of Power Between Hypothalamic Peptidergic Neurons in the Control of Feeding. (2018) Proceedings of the National Academy of Sciences.
- Majumder R., Feola I., Teplenin A.S., de Vries A.A., Panfilov A.V., & Pijnappels D. Optogenetics Enables Real-Time Spatiotemporal Control Over Spiral Wave Dynamics in an Excitable Cardiac System. (2018) eLife.
- Tabor, K.M., Smith, T.S., Brown, M., Bergeron, S.A., Briggman, K.L., & Burgess H.A. Presynaptic Inhibition Selectively Gates Auditory Transmission to the Brainstem Startle Circuit. (2018) Current Biology.
- Virag, T.T., Cserep, C., Schlingloff, D., Posfai, B., Szonyi, A., Sos, K.E., Kornyei, Z., Denes, A., Gulyas, A.I., Freund, T.F., & Nyiri, G. Co-Transmission of Acetlycholine and GABA Regulates Hippocampal States. (2018) Nature Communications.
- Rhomberg, T., Rovira-Esteban, L., Vikor, A., Paradiso, E., Kremser C., Nagy-Pal, P., Papp, O.I., Tasan, R., Erdelyi, F., Szabo, G., Ferraguit, F. & Hajor, N. VIP-Immunoreactive Interneurons Within Circuits of the Mouse Basolateral Amygdala. (2018) Journal of Neuroscience.
- Teplenin, A.S., Dierckx, H., D.A., & Panfilov, A.V. Paradoxical Onset of Arrhythimic Waves from Depolarizes Areas in Cardiac Tissue Due to Curvature-Dependent Instability. (2018) Physical Review X.
- Che, A., Babij, R., Iannone, A.F., Fetcho, R.N., Ferrer, M., Liston, C., Fishell, G., & De Marco Garcia, N.V. Layer I Interneurons Sharpen Sensory Maps during Neonatal Development. (2018) Neuron.
Select illumination of individual cells expressing ChR2 in slice using the Polygon (Courtesy of Matthew Tran & Dr. Blake Richards, McGill University).
Neuroscience Optogenetics
Optogenetics has advanced the field of neuroscience by providing a precise method for manipulating neuronal activity. The ability to perform cellular-resolution optogenetics enables researchers to stimulate select optogenetic-expressing neurons (e.g. ChR2, NpHr, Chrimson etc.) in slice or culture during electrophysiological recordings or calcium imaging to map neural circuitry.
Relevant Publications Using Polygon:
Cell Biology Optogenetics
Optogenetic toolkits have expanded to offer cell biologists unprecedented spatiotemporal molecular and cellular control. From cell migration to gene regulation or developmental processes, cell biologists now have the ability to elicit specific biological responses with an incredible amount of precision in specific cells within organisms or subcellular regions of individual cells using optogenetics.
Select gradient illumination of a cell expressing a novel optogenetic construct using the Polygon (Courtesy of Elliot Dine & Dr. Jared Toettcher, Princeton University).
Relevant Publications Using Polygon:
Johnson HE et al. (2020). Optogenetic Rescue of a Patterning Mutant. Current Biology.
Photoactivation/Photoconversion
Advances in fluorescent protein technology have made photoactivation techniques broadly available in the cell biology lab. By using these techniques, researchers can fluorescently highlight/label specific protein populations in the cell such that their movement and behavior can be tracked over time. Also, photoactivation methods can be applied at different scales, allowing researchers to track proteins within cells, or alternatively to track cells within tissues.
Relevant Publications Using Polygon:
Photopatterning
Many techniques have been developed to manipulate the different properties of the cellular microenviroment. Recently, the advent of light-induced biochemical compounds has allowed scientists to define the biochemical and structural features of cellular matrices by photo micropatterning and photolithography. Scientists can precisely define the spatial distribution of UV light to create unique cellular microenvironments without any use of masks or physical contact with the substrate.

UV Photopatterning (Courtesy of Gayla Berg, University of Colorado).
Relevant Publications Using Polygon:

Optoeletronic tweezers to move specific cells (Courtesy of Dr. Shuailong Zhang, University of Toronto).
Microfluidics
Micro-Fluidic devices are made by photo-patterning thin film precursor materials on a substrate to create microscopic channels, chambers, and even flow-controlling mechanisms. Small amounts of fluids containing reactive chemicals or other substances can then flow through the micro-channels and mix in the micro-chambers of the device and react with other substances that may be deposited in the micro-channels.
Relevant White Papers & Application Notes Using Polygon:
White Paper: Optical Patterned Illumination in Microfluidics
Models
Different Models Optimized for Life Science Applications
Different Models Optimized for Life Science Applications
Mightex offers multiple Polygon models that have been designed with different features to meet the needs of a wide-range of applications that use patterned illumination. Please see below for all available Polygon models.
Polygon1000-G
The Polygon1000-G is a flexible solution for patterned illumination, as this patterned illuminator can be used with any lightsource (350-1000nm) that accepts a 3mm core lightguide. Thus, the Polygon1000-G provides future flexibility for different wavelengths and lightsources, depending on your application.
- Wavelengths: 350-1000nm*
- Lightsource: lightguide-coupled
- Add-on front tube available for large field of view
Key Applications: Neuroscience Optogenetics, Cell Biology Optogenetics, Photoactivation, Photoconversion
* Focus readjustment may be needed when using two wavelengths that are greater than 350nm apart.
Polygon1000-DL
The Polygon1000-DL is a flexible solution for a large field of view or high-power patterned illumination applications, as this patterned illuminator can be used with any fiber-coupled lightsource (350-1000nm), such as high-power lasers. This Polygon1000 model has been designed for high-power applications, such as in vivo optogenetics.
- Wavelengths: 350-1000nm
- Lightsource: fiber-coupled
Key Applications: In Vivo Optogenetics, Cortex-Wide Optogenetics, Photoactivation
* Focus readjustment may be needed when using two wavelengths that are greater than 350nm apart.

Polygon1000-G (Standard)

Polygon1000-G (Large Field-of-View)
Resources
Webinars
White Paper/Applications Notes
Contact Us
600+ Labs Worldwide, 100+ Publications Using Polygon
Mightex’s market-leading Polygon DMD pattern illuminator provides precise spatiotemporal control of light with subcellular resolution, making it the perfect illumination tool for life science research.
Cellular-Resolution Optogenetics and Photostimulation
Simultaneous Multi-Region Illumination
Subcellular Resolution
Compatible with Any Microscope

State-of-the-Art Illumination Technology
The Polygon uses digital mirror device (DMD) technology to illuminate multiple regions simultaneously. A DMD is composed of hundreds of thousands of micro-mirrors that can be individually turned on to reflect light onto the sample. Thus, you can control each mirror to control the area(s) of illumination and create any number of different sized patterns. The Polygon DMD pattern illuminator can be mounted into the infinity-path of any microscope.

Polygon Key Features
Simultaneous Optogenetic Stimulation of Individual Cells or Sub-Cellular Components
Select illumination of individual cells expressing ChR2 in slice using the Polygon (Courtesy of Matthew Tran & Dr. Blake Richards, McGill University).
The Polygon enables scientists to precisely control where light will hit their sample. With subcellular resolution, the Polygon can illuminate any cellular feature in any shape or size simultaneously on their sample.
Stimulation of Any Optogenetics or Photostimulation Probe

The Polygon provides great flexibility when it comes down to wavelength selection. From UV to VIS/NIR range, the Polygon can project light of different colours suitable for your light-sensitive constructs.
Spatio-Temporal and Intensity Control of Light
Select gradient illumination of a cell expressing a novel optogenetic construct using the Polygon (Courtesy of Elliot Dine & Dr. Jared Toettcher, Princeton University).
Control the initiation, duration, and intensity of light stimulation patterns using the Polygon. Create different waveforms to control the light intensity and duration outputted from the Polygon.
In Vitro and In Vivo Optogenetic Stimulation
The Polygon can be used with a wide-range light sources, including LEDs and lasers. The Polygon is also compatible with Mightex’s OASIS Implant for freely-behaving imaging and optogenetics and OASIS Macro for cortex-wide imaging and optogenetics.
UV Micropatterning with Sub-Micron Resolution

UV Photopatterning (Courtesy of Gayla Berg, University of Colorado).
The Polygon is compatible with UV light sources (350nm+) and provides sub-micron illumination for photopatterning and microfabrication experiments.
Integrates into Any Electrophysiology, Two-Photon or Confocal Microscope Setup
Illumination patterns using the Polygon synchronized with slice electrophysiological recordings (Courtesy of Matthew Tran and Dr. Blake Richards, McGill University).
The Polygon is designed to be coupled into the infinity space of any microscope model (Leica, Nikon, Olympus, Zeiss) with Mightex’s microscope-specific adapters. The Polygon comes with BNC connectorized TTL trigger input and output for easy synchronization with different lab equipment.
Powerful User-Friendly Software for Life Science Experiments
Mightex’s PolyScan software platform is bundled with every Polygon to help you execute sophisticated patterned illumination experiments for your research.
600+ Worldwide Polygon Customers
100+ Publications Using the Polygon
Contact Information
- (In Canada) 200 Consumers Road, Suite 805,
Toronto, Ontario M2J 4R4, CANADA - (Canada) 1-416-840-6115
- 1-416-840 6541
- (In US) 1241 Quarry Lane, Suite 105,
Pleasanton, CA 94566, USA - (US) 1-925-218-1885
- 1-416-840 6541
Follow Us On Social Media
Company Policies
Subscribe To Mailing List
















