Voltage imaging enables scientists to visualize neural activity using fluorescent activity sensors that are sensitive to changes in voltage
This guide explores questions related to voltage imaging, including what is voltage imaging? What equipment is necessary for voltage imaging?
Electrophysiology is the golden standard for recording individual neuron activity. By recording electrical changes across the cell membrane, electrophysiology is able to detect action potentials in real-time (Petersen, 2017). Due to the complex nature of the technique, however, electrophysiology cannot be used to measure the activity of hundreds of neurons simultaneously, although such capability is essential for understanding the complex interactions between populations of neurons leading to cognition, behaviour, and sensory processing.
Calcium imaging has helped scientists overcome this limitation by enabling them to visualize the activity of large neuron populations. Using genetically-encoded calcium indicators, scientists are able to measure neural activity changes in hundreds of neurons simultaneously through fluctuations in fluorescence signals (Grienberger & Konnerth, 2012). Although calcium imaging has enabled the ability to visualize large neural population activity, it relies upon an indirect indicator of neural activity (i.e.calcium influx), which is inherently very slow, and hence, it is unable to faithfully reflect the relatively fast neuron activity (Grienberger & Konnerth, 2012). Electrophysiology, on the other hand, is much faster and it is able to reflect real-time changes in neural activity.
Is there a technique that can combine the strengths of both electrophysiology and calcium imaging to measure neural activity?
Voltage Imaging
What is Voltage Imaging?
Building upon the strengths of electrophysiology and calcium imaging, voltage imaging enables scientists to visualize neural activity using fluorescent activity sensors that are sensitive to changes in voltage (Knopfel & Song, 2019). Unlike calcium imaging that relies on an indirect indicator, voltage imaging reflects real-time neuronal firing, and thus, it is fast enough to detect action potentials and subthreshold responses (Knopfel & Song, 2019). At the same time, voltage Imaging enables scientists to visualize a large number of neurons simultaneously.
The development of genetically encoded voltage indicators (GEVIs) has enabled neuroscientists to study specific cell-types (e.g. excitatory or inhibitory neurons). Some of the recently developed GEVI’s include ASAP3, Voltron, QuasAR3, and SomArchon (Villette et al., 2019; Abdelfattah et al., 2019; Adam et al., 2019; Piatkevich et al., 2019). However, since the field of voltage imaging is quite new, the GEVI toolbox is constantly growing with new indicators optimized for certain experiments, such as two-photon imaging.
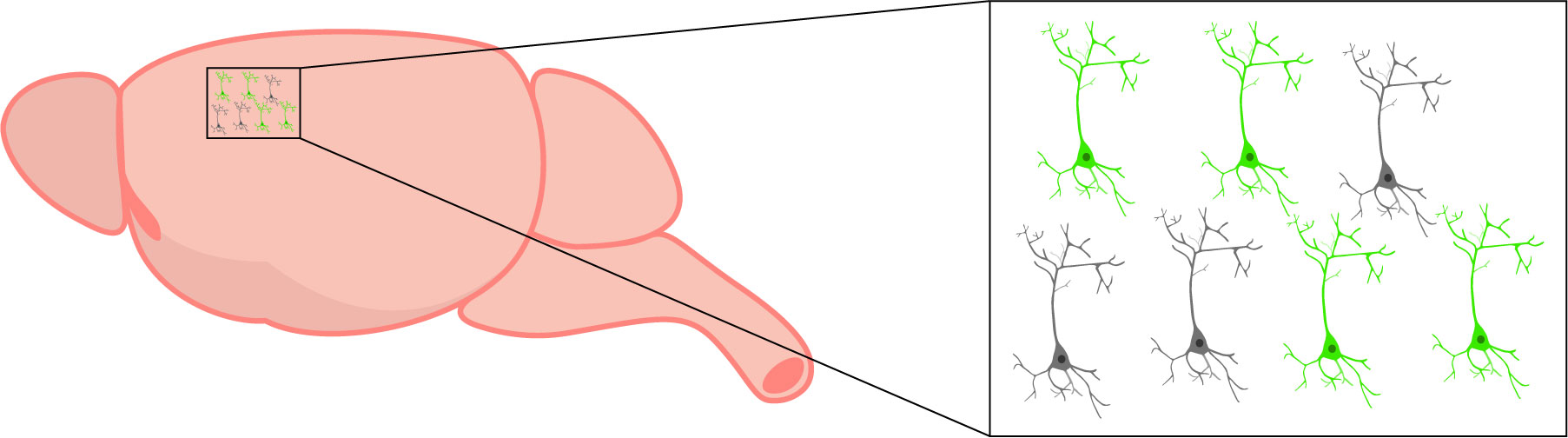
Voltage imaging sensors are expressed in specific cells (in green). Changes in activity are only detected in cells expressing the sensor.
Why Use Voltage Imaging?
Voltage imaging, being a relatively new technique, is employed by only a few labs and currently being optimized for more global use. The potential of voltage imaging’s capabilities can help advance the ability of tracking real-time neural activity in large neuron populations.
Voltage imaging visualizes a direct indicator of activity by measuring changes in voltage, similar to electrophysiology. Measuring electrical changes, or action potentials, provides a method to neural activity changes in real-time with high speed, reflecting the key marker of neural activity (Knopfel & Song, 2019). This is in contrast to calcium imaging which measures the indirect and slow indicator of calcium influx (Grienberger & Konnerth, 2012).
GEVIs enable the visualization of specific neural populations, similar to calcium imaging. Compared to electrophysiology that can measure only a few neurons simultaneously, voltage imaging can identify and measure the activity of multiple neurons simultaneously to track activity long-term.
Voltage imaging has the ability to measure neural activity, reflecting electrical changes, in behaving animals. This provides scientists with the highest temporal and spatial precision to track neural activity over time in multiple neurons to correlate neuron firing, behaviour, and cognition. Electrophysiology can be performed in vivo; however, it cannot be used to spatial track large populations of neurons in vivo, making voltage imaging an essential technique for recording neural activity in vivo.
Due to the nature of an early-stage technique, voltage imaging has a few challenges associated with both the biology and equipment, which need to be properly addressed as the technology continues to evolve.
Voltage indicators are fast, reflecting millisecond changes in neural activity, and, as a result, voltage imaging requires a high frame-rate scientific camera (Knopfel & Song, 2019; Yang & St. Pierre, 2016). Some cameras currently available in the market are able to reach high enough frame rates, but they may require cropping the field of view to reach the adequate speed, thereby limiting the field of view to a smaller number of neurons (Knopfel & Song, 2019; Yang & St. Pierre, 2016).
In addition, a fast frame rate also leads to shorter exposure time, meaning that a camera used for voltage imaging must also have high sensitivity, in order to capture significant signals from GEVIs (Knopfel & Song, 2019; Yang & St. Pierre, 2016).
Furthermore, stronger light sources can help reach the required light intensity; however, higher light intensity on the sample increases the likelihood of photobleaching the sample, decreasing the length of the experiment (Knopfel & Song, 2019; Yang & St. Pierre, 2016). Some research labs are optimizing the brightness of GEVis to help resolve this problem. Through the development of calcium imaging, there has been a significant increase in brightness of light sources, so given time these problems should be properly addressed.
Another issue with voltage imaging is low signal-to-noise ratio. This is associated with high light scattering and relatively low expression in tissue (Knopfel & Song, 2019; Yang & St. Pierre, 2016). Scientists are also working on optimizing GEVIs to improve the signal-to-noise ratio.
To summarize, while calcium imaging is currently the most well-established technique for visualizing large neural circuits in vivo, voltage imaging may become the future of imaging neural activity, thanks to its superior real-time accuracy. Further development of brighter, more sensitive, better signal-to-noise, and better expression will help make voltage imaging a powerful tool.
If someone were to describe voltage imaging to you, it might sound quite simple.
When you dig deeper, you start to appreciate the complexity associated with the biology and equipment to perform voltage imaging.
You’re probably asking, what are the necessary components to perform voltage imaging?
Biological Components
Brains don’t naturally express genetically encoded voltage indicators (GEVIs), meaning there are biological steps required in order to perform voltage imaging in animals. First, you must express the genetic indicator in the brain; and second, depending on if you are performing in vivo experiments, you need to implant an imaging probe to collect fluorescent signals from the brain.
Genetic Sensor Expression
The first, and most important step, is achieving optimal GEVI expression in your animal model.
Mice are the most common animal model used for voltage imaging due to the advancement of genetic mice models (Daigle et al. 2018) and viral expression in the brain.
Viral expression involves injecting a virus encoding a GEVI in the brain. This virus is linked to a gene of interest to target expression in a specific cell-type.
A crucial step associated with viral expression is testing varying dilutions of the virus to obtain optimal expression in the brain (Resendez et al. 2016). Too little expression can lead to no signal, and over-expression can lead to high fluorescence background noise – or even cell death!
Neuroscientists employ viral expression to regulate GEVI expression. This is useful because expression can vary depending on the brain region, cell-type, or virus. In addition, neuroscientists can use viral expression to express GEVIs in brain projections to map neural circuits across brain regions.
Imaging Probe (For In Vivo Experiments)
After successful GEVI expression, you need to access the fluorescent signal inside the brain for in vivo experiments. But how can you see into the brain when it’s covered by both skin and skull?
Generally, this involves surgically implanting an imaging probe into the brain where the GEVI is expressed. There are three types of probes (optical cannula, cortical window, GRIN lens) that are used for voltage imaging.
Optical cannulas enable light to be delivered to and collected from the brain. These probes are used in Fiber Photometry experiments. Due to their design, optical cannulas are only capable of collecting a population signal, providing little or no spatial resolution to distinguish individual cells.
In contrast, cortical windows replace a large portion of the skull with a glass window. Neuroscientists employ cortical windows when imaging a large cortical region on the surface of the brain. Combined with a camera, cortical windows provide access to the cortex for single-cell resolution recordings.
Lastly, a GRIN lens is a microendoscopic probe that can be implanted in the brain to image deep regions of the brain (up to 8mm) with single-cell resolution. GRIN lenses differ in lengths, enabling neuroscientists to image both shallow and deep brain regions. To minimize tissue damage, scientists usually use GRIN lenses with relatively small diameters (e.g. 0.5mm ~ 1mm). As a result, imaging with GRIN lenses usually provides a relatively small field of view, especially compared to cortical windows.
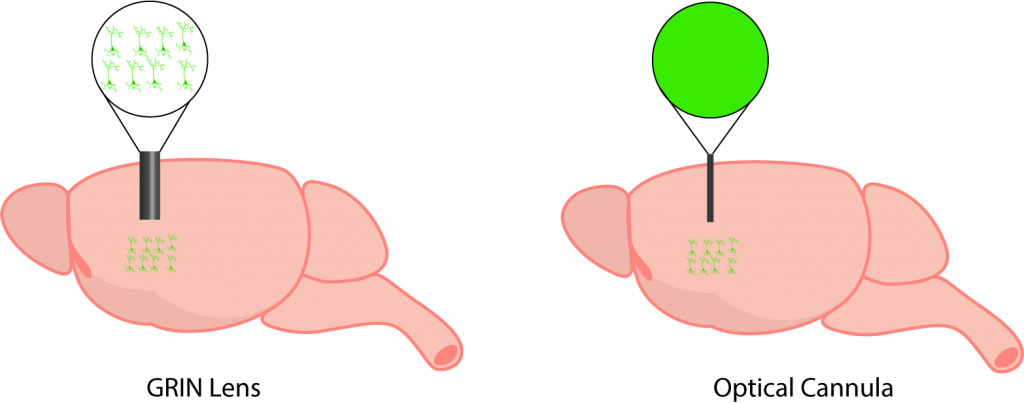
Comparison between imaging with a GRIN lens and optical cannula.
Equipment Components
Now that you have all biological components setup, including optimal GEVI expression and imaging probe implantation, if necessary, you need equipment to record fluorescent signals.
These are the main components required for voltage imaging:
Imaging System
You will select your imaging system based on the sample being studied. There are pros and cons to each, which is related to the biological and technological constraints, but each system is designed for a specific application. Here are four potential imaging systems that can be used for voltage imaging:
- One-photon microscope for slice imaging, such as Mightex’s OASIS Micro.
- Two-photon microscope for in vivo head-fixed experiments
- Mesoscope for in vivo head-fixed large field of view experiments, such as Mightex’s OASIS Macro
- Optical fiberscope for in vivo freely-behaving experiments, such as Mightex’s OASIS Implant.
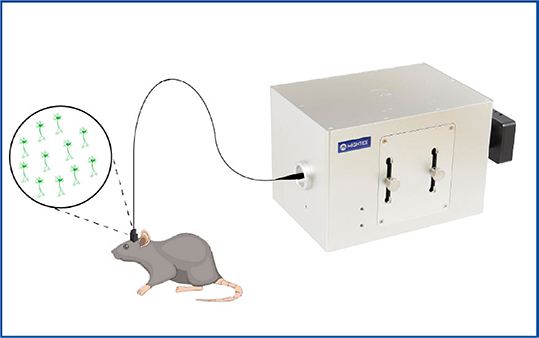

Mightex’s OASIS Implant is a system for freely-behaving or head-fixed voltage imaging.
Imaging Device
You need to detect and analyze fluorescent signals from the brain. This is made possible using an imaging device. Three types of imaging devices are used for in vivo imaging systems: 1) scientific camera, 2) PMT, and 3) photodetector. Which imaging device is used is somewhat dependent on the voltage imaging application.
If you’re using a scientific camera for voltage imaging, it is necessary to select a camera with a high-frame rate to capture signal fast enough and high-sensitivity to detect weak signals that may come from voltage imaging (Knopfel & Song, 2019; Yang & St. Pierre, 2016).
Successful voltage imaging is a balancing act between biology and equipment. Luckily, both the biology and equipment are constantly being optimized for better performance and ease of use.
Light Source and Filter Set
You need to illuminate and collect fluorescent signals from the brain to perform voltage imaging. GEVIs function by generating fluorescence signals, such that they have an excitation and emission spectrum (Quicke et al., 2019). To excite GEVIs and collect the emitting signal you require two components: a light source and dichroics/filters.
For excitation of the GEVI, LED light sources or lasers may be employed, depending on the experiment. For example, voltage imaging signal may require high optical power to detect it, so it is important to select a light source that has the appropriate output for the probe you are studying (Knopfel & Song, 2019; Yang & St. Pierre, 2016).
Importantly, the correct excitation wavelength must be selected, and this is where the second component is necessary: dichroics and filters allow proper transmission of the correct excitation wavelength and transmission of the correct emission signal to the imaging device.
DMD (Digital Micromirror Device)
A DMD (digital micromirror device), such as Mightex’s Polygon DMD Pattern Illuminator, enables scientists to control precisely which areas on the sample the light will illuminate. The problem with using a standard wide-field light source for voltage imaging is it illuminates the entire sample, and this may lead to strong fluorescence background noise and fast photobleaching of the sample, affecting the signal quality and length of the experiment (Adam et al., 2019).
In order to avoid these problems, scientists have recently employed DMDs to focus light onto the cells they are studying to prevent these problems. With a DMD to focus the light, you can improve signal quality by reducing background fluorescence and reduce photobleaching by only direct light to the specific areas of interest on the sample (Adam et al., 2019). As a result, DMD pattern illuminators, such as Mightex’s Polygon, may become an important technology to overcome the biological limitations of GEVIs.
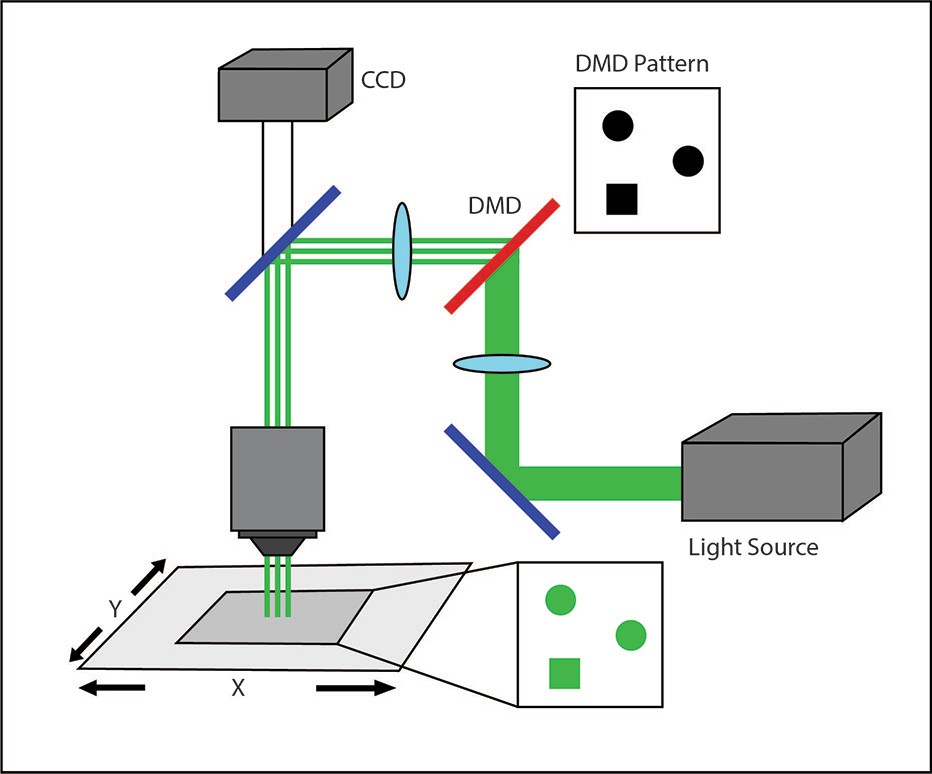
How a DMD setup controls illumination on the sample.
The invention of optogenetics has enabled scientists to use light to turn cell-type specific neural activity on or off with millisecond-precision to probe neural circuit function (Yizhar et al. 2011). Complementing optogenetics, voltage imaging enables scientists to observe cell-type specific neural circuit activity with cellular-resolution using fluorescent indicators (Knopfel & Song, 2019).
With voltage imaging, scientists can closely replicate electrophysiology measurements, but with the advantage of visualizing large neural populations. Integrating optogenetics with voltage imaging opens the possibility of performing all-optical electrophysiology (Hochbaum et al. 2014).
All-optical electrophysiology would enable scientists to image the activity of large neuron populations and simultaneously perturb cellular activity using optogenetics to both ‘read’ and ‘write’ neural activity. The integration of optogenetics and voltage imaging opens the door to these techniques being performed simultaneously in the same experiment to investigate the causal link between neural circuit activity, function, and behaviour.
To perform all-optical electrophysiology experiments with both voltage imaging and optogenetics, scientists require optimized biology and the appropriate equipment.
All-Optical Biology
A wide-range of optogenetic probes have been developed, differing in function (excitation or inhibition), activation time, and expression properties.
Optogenetic probes are excited by wavelengths of light ranging from blue to red, depending on their properties, and they have only an excitation spectrum to activate their excitation/inhibitory properties (Yizhar et al. 2011).
In contrast, GEVIs behave in the same manner as a fluorophore (e.g. GFP), such that they exhibit excitation and emission spectra. However, the fluorescent signals of GEVIs are dependent on changes in voltage (i.e. increased fluorescence emission) and display dynamic behaviour, unlike a static signal emitted from a fluorophore (Knopfel & Song, 2019).
One problem encountered by combining optogenetics and voltage imaging is overlapping excitation spectra, such that both probes have a similar excitation wavelength. This means that, when trying to image the GEVI, you can potentially activate the optogenetic probe at the same time. Thus, due to potential optical crosstalk, it is not possible to determine if measured changes in the voltage imaging signal are due to natural changes in neural activity or optogenetic-induced changes.
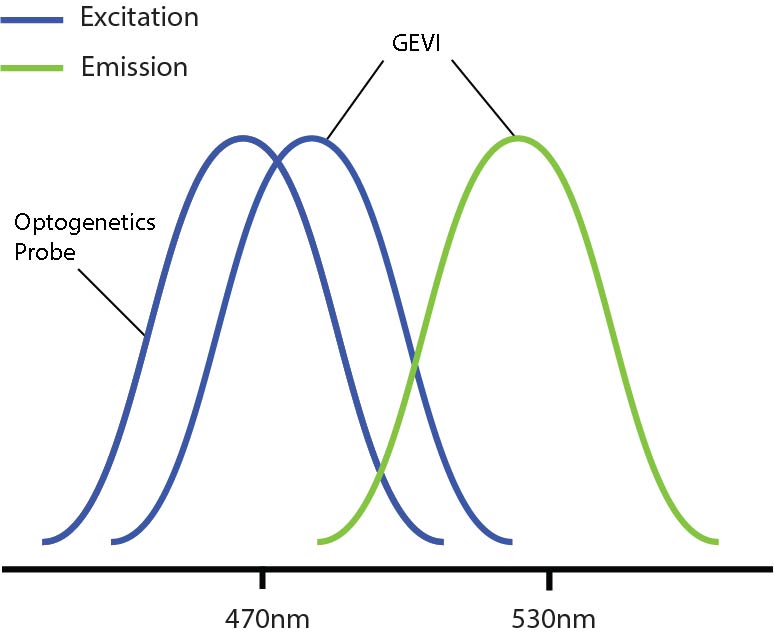
Overlapping excitation spectra of the optogenetics probe and voltage imaging probe.
Optical crosstalk in all-optical electrophysiology experiments can be reduced by selecting imaging indicators and optogenetic probes with non-overlapping excitation spectrums (Emiliani et al. 2015).
An example of this is used by Fan et al. 2020 where they used red-shifted GEVI SomArchon and blue-shifted optogenetic probe CheRiff for all-optical electrophysiology. This decreased the likelihood of crosstalk by reducing or even preventing indirect activation of the optogenetic construct during imaging.
The further optimization of voltage imaging and optogenetics probes (e.g. sensitivity, excitation spectra) will help prevent optical crosstalk in all-optical electrophysiology experiments.
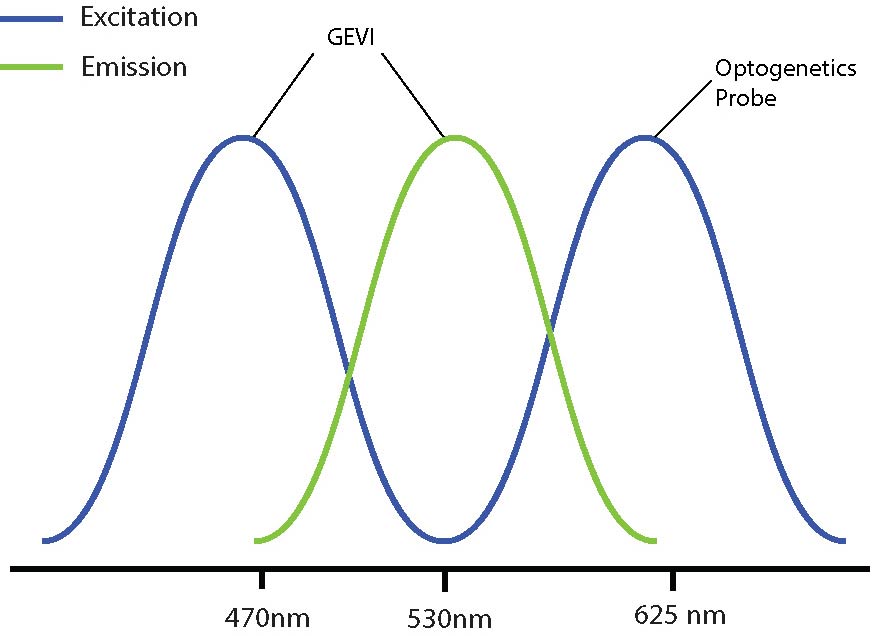
By selecting a wavelength shifted opsin, you can reduce any potential optical crosstalk.
Equipment: Widefield vs. Cellular-Resolution Optogenetics
Once the biology is optimized, you will require the appropriate equipment. For all-optical electrophysiology experiments, there are two options for optogenetics: widefield and cellular-resolution.
Widefield optogenetics uses a standard light source to illuminate the entire field of view. Thus, all neurons will be stimulated indiscriminately, with no ability to select individual neurons.
In comparison, the ability to illuminate select neurons within a population of optogenetic-expressing neurons is called cellular-resolution optogenetics (Shemesh et al. 2017). This method requires more sophisticated spatial specificity to selectively stimulate individual neurons within an optogenetic-expressing population.
The ability to illuminate certain individual neurons is not a limitation of optogenetics per say, but a limitation of the standard optogenetic light sources used in most optogenetic systems. As mentioned above, standard light sources can only illuminate all cells within the field of view, and they have no ability to control where they illuminate to target individual neurons.
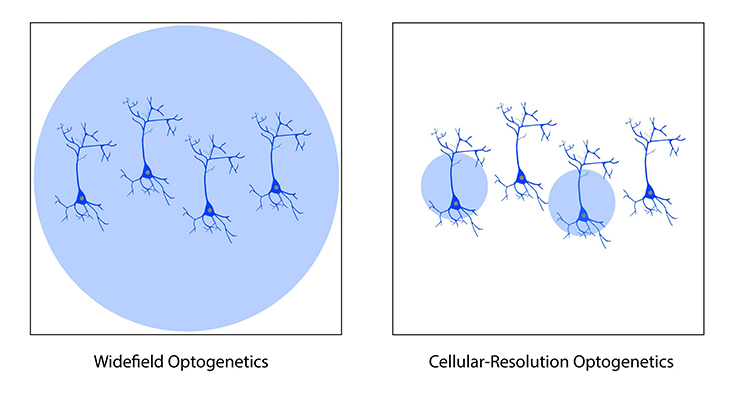
Comparison between widefield and cellular-resolution optogenetic stimulation.
The development of spatially targeted light technologies have enabled researchers to control where the light illuminates the sample, such as an individual cell, making cellular-resolution optogenetics possible (Ronzitti et al. 2017). For example, a digital micromirror device (DMD), such as Mightex’s Polygon Patterned Illuminator, allows researchers to illuminate multiple select individual neurons or regions simultaneously to perform cellular-resolution optogenetics.
Cellular-resolution optogenetics has multiple applications in the field of neuroscience. For example, scientists can study neural circuits and decode neural patterns at the level of individual neurons (Anastasiades et al. 2020; Tran et al. 2019). These types of studies have been carried out both in vitro and in vivo using different technologies (Shemesh et al. 2017; Anastasiades et al. 2020; Tran et al. 2019; Chen et al. 2019). The experimental applications for cellular-resolution optogenetics is endless.
Depending on your all-optical electrophysiology experiment, there are options available for both widefield and cellular-resolution optogenetics integration.