Patterned illumination technologies enable researchers to selectively illuminate parts of their sample for optogenetic stimulation. One application of this is cellular-resolution optogenetics: the ability to selectively illuminate individual neurons within an optogenetic-expressing population (to further understand the difference between widefield and cellular-resolution optogenetics, see this blog post). Beyond this application, patterned illumination can enable researchers to perform more precise and intricate optogenetic experiments.
How can patterned illumination be used in neuroscience optogenetics experiments?
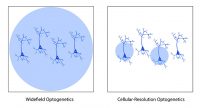
Channelrhodopsin-Assisted Circuit Mapping (CRACM) – Kissinger et al. 2020 Cell Reports
Kissinger et al. 2020 published a paper in Cell Reports examining perceptual deficits in a mouse model of Fragile X syndrome (Fmr1 knockout mice). Previously, their lab showed that visually-evoked low-frequency oscillations occur in V1 during encoding of a familiar stimulus. Kissinger et al. 2020 sought to examine any potential abnormalities in Fmr1 knockout mice low-frequency oscillations during perception and associated circuit changes.
Using local field potentials and single-unit recordings, the authors demonstrated that following the presentation of a familiar visual stimulus Fmr1 mice exhibited shorter duration, lower power, and lower frequency of oscillations in multiple layers of visual cortex.
Kissinger et al. 2020 followed this by investigating any potential connectivity changes in V1 by performing channelrhodopsin-assisted circuit mapping (CRACM) using Mightex’s Polygon DMD Illuminator. Using this technique, the authors were able to examine
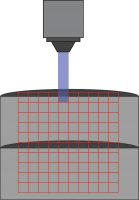
Mightex’s Polygon DMD Illuminator scans across a 10 x 10 grid to stimulate individuals areas during a patch-clamp recording.
single-cell connections between cells using optogenetics.
Channelrhodopsin was expressed in control and Fmr1 knockout animals by crossing them with Thy1-ChR2-YFP mice. Naive or experienced animals were sacrificed, and patch-clamp electrophysiology was used to test connections between cells. Specifically, connections between layer 5 – layer 4 and layer 5 – layer 5 cells were tested. While cells were patched, individual cells were stimulated in a 10 by 10 grid (0.67 mm x 0.67 mm) using the Polygon (covering a square from L2/3 to L5). A 470 nm LED light source with 10 ms optogenetic light pulses through a 10x objective was used (0.3 mW under the objective) to stimulate individual neurons.
Kissinger et al. showed a significant potentiation in layer 5 to layer 4 fast-spiking cells in control animals, but no change in Fmr1 knockout animals following perceptual experience. In addition, a significant depression in layer 5 to layer 5 neurons was seen in Fmr1 knockout animals following perceptual experience.
This study demonstrates significant differences in how an animal model of Fragile X syndrome encodes information. As well, using Mightex’s Polygon for CRACM to demonstrate associated circuit changes in Fmr1 knockout mice.
Focal Optogenetic Stimulation in the Cerebellum – Gruver & Watt 2019 Frontiers in Synaptic Neuroscience
Gruver & Watt 2019 published a paper in Frontiers in Synaptic Neuroscience looking to understand the parameters for focal optogenetic stimulation of purkinje cell somas and axons in the cerebellum using Mightex’ Polygon DMD Illuminator. In
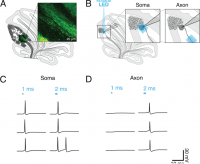
(A) Schematic of sagittal cerebellar slice. Inset is a maximal intensity projection of abtwo-photon stack showing ChR2(H134R)-EYFP expression (green) in axons in the white matter. (B) Recording configuration. Insets show the focal region of photostimulation (blue square) and somatic recording electrode. (C and D) Representative current-clamp traces of optically-evoked action potentials evoked following somatic (C) and axonal (D) stimulation. Blue bars above trace indicate onset and duration of light pulse.
addition, they wanted to understand the connections between Purkinje and DCN neurons in the cerebellum using focal optogenetic stimulation.
Cerebellum slices expressed ChR2 and neuronal activity was measured using patch-clamp electrophysiology. While the soma of a cell was patched, the soma or white matter (axon) of that cell was individually illuminated using Mightex’s Polygon DMD illuminator with a 40 um x 40 um square. Focal optogenetic stimulation was performed under a 60x objective with an approximate 100 mw/mm2 intensity at the focal plane.
Gruver & Watt 2019 tested different pulse parameters to determine the ideal optogenetic stimulation to elicit a single action-potential. Longer optogenetic stimulation pulses were required for axons (2-3 ms) compared to soma (1 ms) to elicit a single action potential.
The authors also tested the connections between Purkinje cells and postsynaptic DCN neurons in the cerebellum using focal optogenetic stimulation. The soma of DCN cells was patched and the axon of the presynaptic purkinje cells was focally stimulated (200 um from DCN cells). Gruver & Watt 2019 determined as optogenetic stimulation pulse duration increased, the post-synaptic response in DCN cells increased.
Gruver & Watt 2019 demonstrate a protocol of successful focal stimulation using Mightex’s Polygon DMD illuminator to elicit single action potentials to test neural circuitry in the cerebellum.
References
- Kissinger ST et al. (2020). Visual Experience-Dependent Oscillations and Underlying Circuit Connectivity Changes are Impared in Fmr1 KO Mice. Cell Reports, 31(1), 107486.
- Gruver KM & Watt AJ (2019). Optimizing Activation of Purkinje Cell Axons to Investigate the Purkinje Cell – DCN Synapse. Frontiers in Synaptic Neuroscience, 11(31).
Next Post
How Can Patterned Illumination be Used in Optogenetics Experiments? (Part 2)




